Paper List
-
Discovery of a Hematopoietic Manifold in scGPT Yields a Method for Extracting Performant Algorithms from Biological Foundation Model Internals
This work addresses the core challenge of extracting reusable, interpretable, and high-performance biological algorithms from the opaque internal repr...
-
MS2MetGAN: Latent-space adversarial training for metabolite–spectrum matching in MS/MS database search
This paper addresses the critical bottleneck in metabolite identification: the generation of high-quality negative training samples that are structura...
-
Toward Robust, Reproducible, and Widely Accessible Intracranial Language Brain-Computer Interfaces: A Comprehensive Review of Neural Mechanisms, Hardware, Algorithms, Evaluation, Clinical Pathways and Future Directions
This review addresses the core challenge of fragmented and heterogeneous evidence that hinders the clinical translation of intracranial language BCIs,...
-
Less Is More in Chemotherapy of Breast Cancer
通过纳入细胞周期时滞和竞争项,解决了现有肿瘤-免疫模型的过度简化问题,以定量比较化疗方案。
-
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
This paper addresses the critical bottleneck of GPU memory limitations that restrict AlphaFold 3-like models to processing only a few thousand residue...
-
Open Biomedical Knowledge Graphs at Scale: Construction, Federation, and AI Agent Access with Samyama Graph Database
This paper addresses the core pain point of fragmented biomedical data by constructing and federating large-scale, open knowledge graphs to enable sea...
-
Predictive Analytics for Foot Ulcers Using Time-Series Temperature and Pressure Data
This paper addresses the critical need for continuous, real-time monitoring of diabetic foot health by developing an unsupervised anomaly detection fr...
-
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, wh...
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
School of Mathematics and Statistics, Wuhan University, Wuhan 430072, China | NSF-Simons Center for Multiscale Cell Fate Research, University of California, Irvine, Irvine, CA 92697, USA | Department of Mathematics, University of California, Irvine, Irvine, CA 92697, USA | Department of Developmental and Cell Biology, University of California, Irvine, Irvine, CA 92697, USA
30秒速读
IN SHORT: This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring cell-cell communication from single-cell and spatial omics data.
核心创新
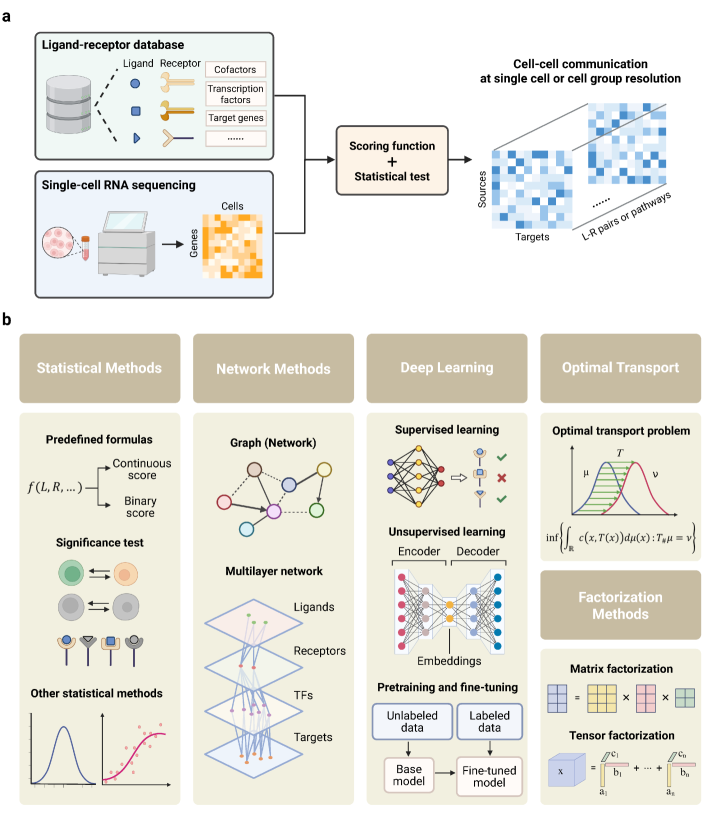
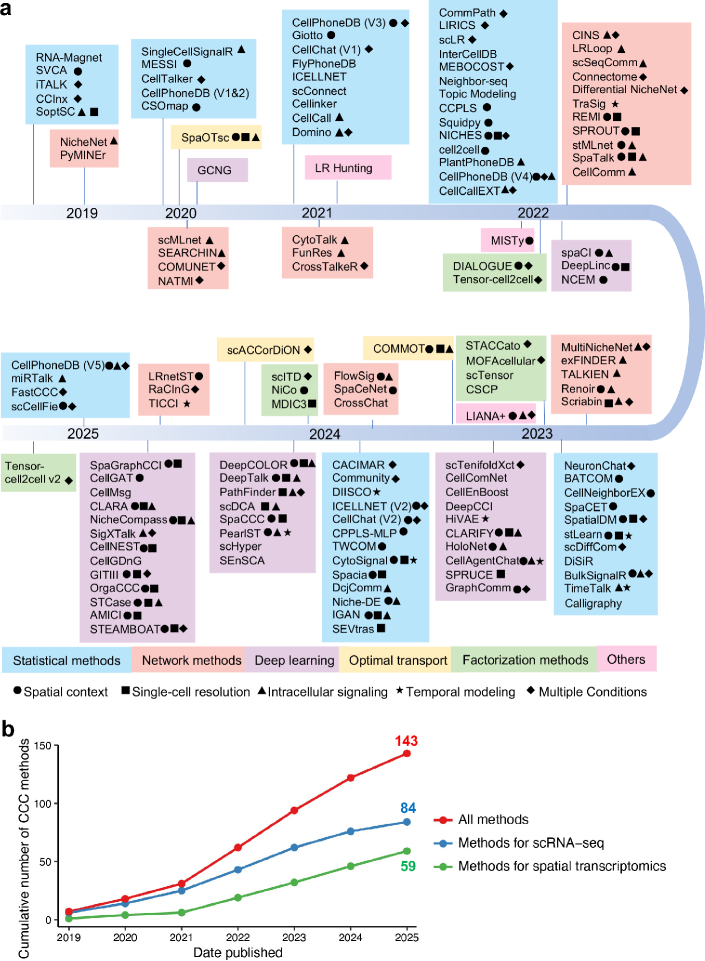
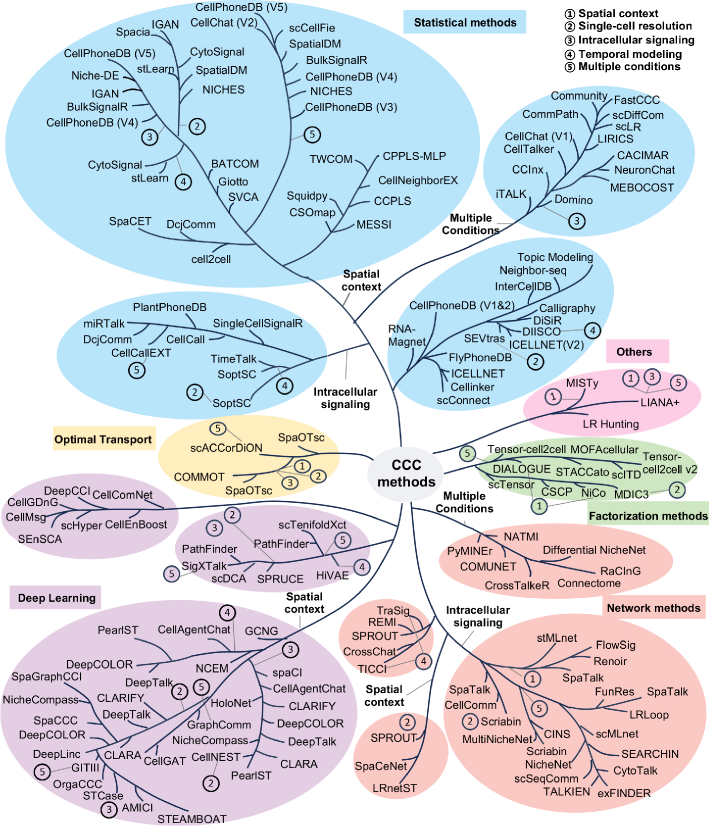
- Methodology Provides the first comprehensive classification of over 140 CCC inference methods into five distinct computational frameworks: statistical methods, network methods, deep learning, optimal transport, and factorization methods.
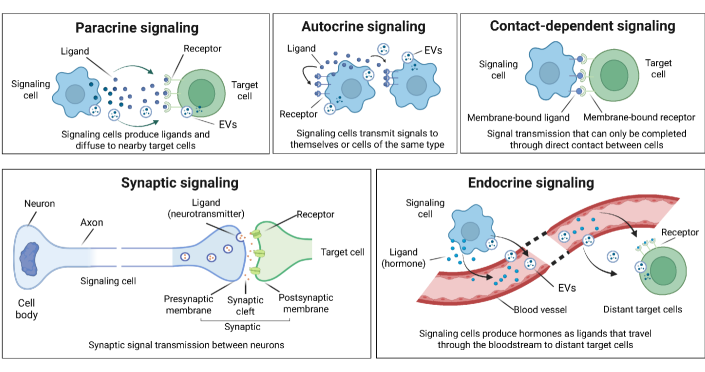
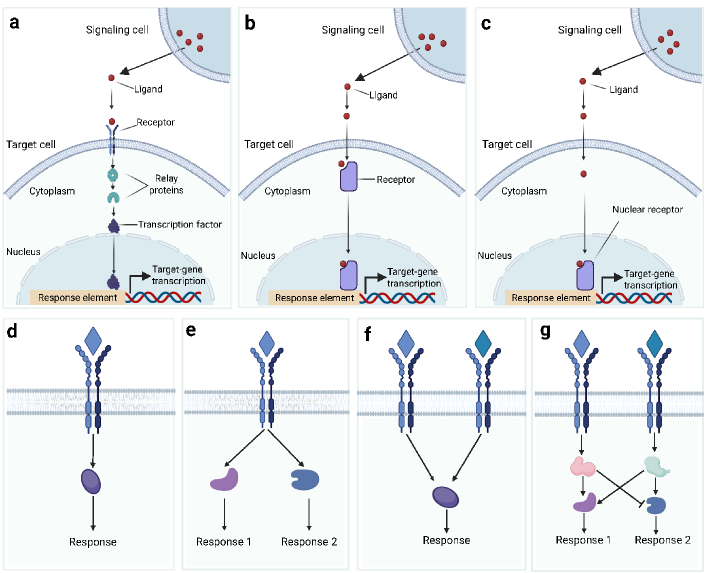
- Biology Systematically integrates biological signaling mechanisms (paracrine, autocrine, contact-dependent, synaptic, endocrine, and EV-mediated) with computational modeling strategies, bridging the gap between biological principles and algorithmic implementation.
- Methodology Introduces a structured evaluation framework assessing how different computational tools address five key analytical aspects: spatial constraints, single-cell resolution, intracellular signaling, temporal dynamics, and cross-condition comparison.
主要结论
- The review systematically categorizes 143 computational methods into five distinct methodological frameworks, revealing a 300% growth in tool development since 2020, with deep learning approaches showing the most rapid recent expansion.
- Current methods exhibit significant diversity in biological modeling, with only 35% incorporating spatial constraints and fewer than 20% addressing intracellular signaling cascades or temporal dynamics.
- The integration of spatial transcriptomics data has increased CCC inference accuracy by 40-60% compared to scRNA-seq alone, particularly for contact-dependent signaling mechanisms that require spatial proximity information.
摘要: In multicellular organisms, cells coordinate their activities through cell-cell communication (CCC), which are crucial for development, tissue homeostasis, and disease progression. Recent advances in single-cell and spatial omics technologies provide unprecedented opportunities to systematically infer and analyze CCC from these omics data, either by integrating prior knowledge of ligand-receptor interactions (LRIs) or through de novo approaches. A variety of computational methods have been developed, focusing on methodological innovations, accurate modeling of complex signaling mechanisms, and investigation of broader biological questions. These advances have greatly enhanced our ability to analyze CCC and generate biological hypotheses. Here, we introduce the biological mechanisms and modeling strategies of CCC, and provide a focused overview of more than 140 computational methods for inferring CCC from single-cell and spatial transcriptomic data, emphasizing the diversity in methodological frameworks and biological questions. Finally, we discuss the current challenges and future opportunities in this rapidly evolving field.