Paper List
-
The Effective Reproduction Number in the Kermack-McKendrick model with age of infection and reinfection
This paper addresses the challenge of accurately estimating the time-varying effective reproduction number ℛ(t) in epidemics by incorporating two crit...
-
Covering Relations in the Poset of Combinatorial Neural Codes
This work addresses the core challenge of navigating the complex poset structure of neural codes to systematically test the conjecture linking convex ...
-
Collective adsorption of pheromones at the water-air interface
This paper addresses the core challenge of understanding how amphiphilic pheromones, previously assumed to be transported in the gas phase, can be sta...
-
pHapCompass: Probabilistic Assembly and Uncertainty Quantification of Polyploid Haplotype Phase
This paper addresses the core challenge of accurately assembling polyploid haplotypes from sequencing data, where read assignment ambiguity and an exp...
-
Setting up for failure: automatic discovery of the neural mechanisms of cognitive errors
This paper addresses the core challenge of automating the discovery of biologically plausible recurrent neural network (RNN) dynamics that can replica...
-
Influence of Object Affordance on Action Language Understanding: Evidence from Dynamic Causal Modeling Analysis
This study addresses the core challenge of moving beyond correlational evidence to establish the *causal direction* and *temporal dynamics* of how obj...
-
Revealing stimulus-dependent dynamics through statistical complexity
This paper addresses the core challenge of detecting stimulus-specific patterns in neural population dynamics that remain hidden to traditional variab...
-
Exactly Solvable Population Model with Square-Root Growth Noise and Cell-Size Regulation
This paper addresses the fundamental gap in understanding how microscopic growth fluctuations, specifically those with size-dependent (square-root) no...
Beyond Bayesian Inference: The Correlation Integral Likelihood Framework and Gradient Flow Methods for Deterministic Sampling
Institute of Mathematics, Polish Academy of Sciences | Interdisciplinary Centre for Mathematical and Computational Modelling, University of Warsaw | Institute for Mathematics, Heidelberg University
30秒速读
IN SHORT: This paper addresses the core challenge of calibrating complex biological models (e.g., PDEs, agent-based models) with incomplete, noisy, or heterogeneous data, where traditional pointwise comparison methods fail due to system sensitivity and intrinsic variability.
核心创新
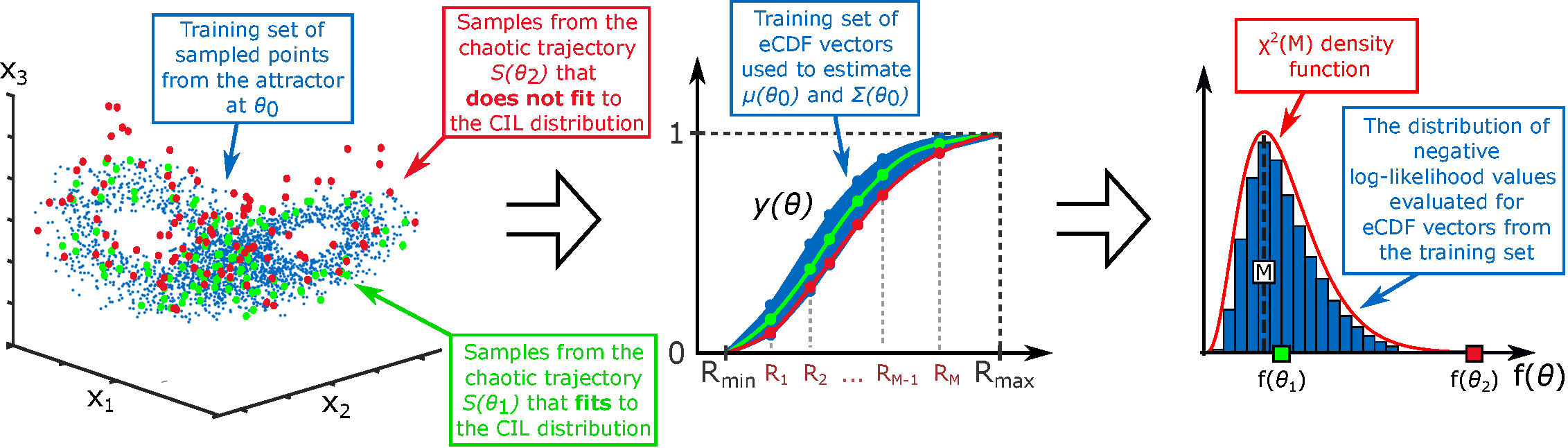
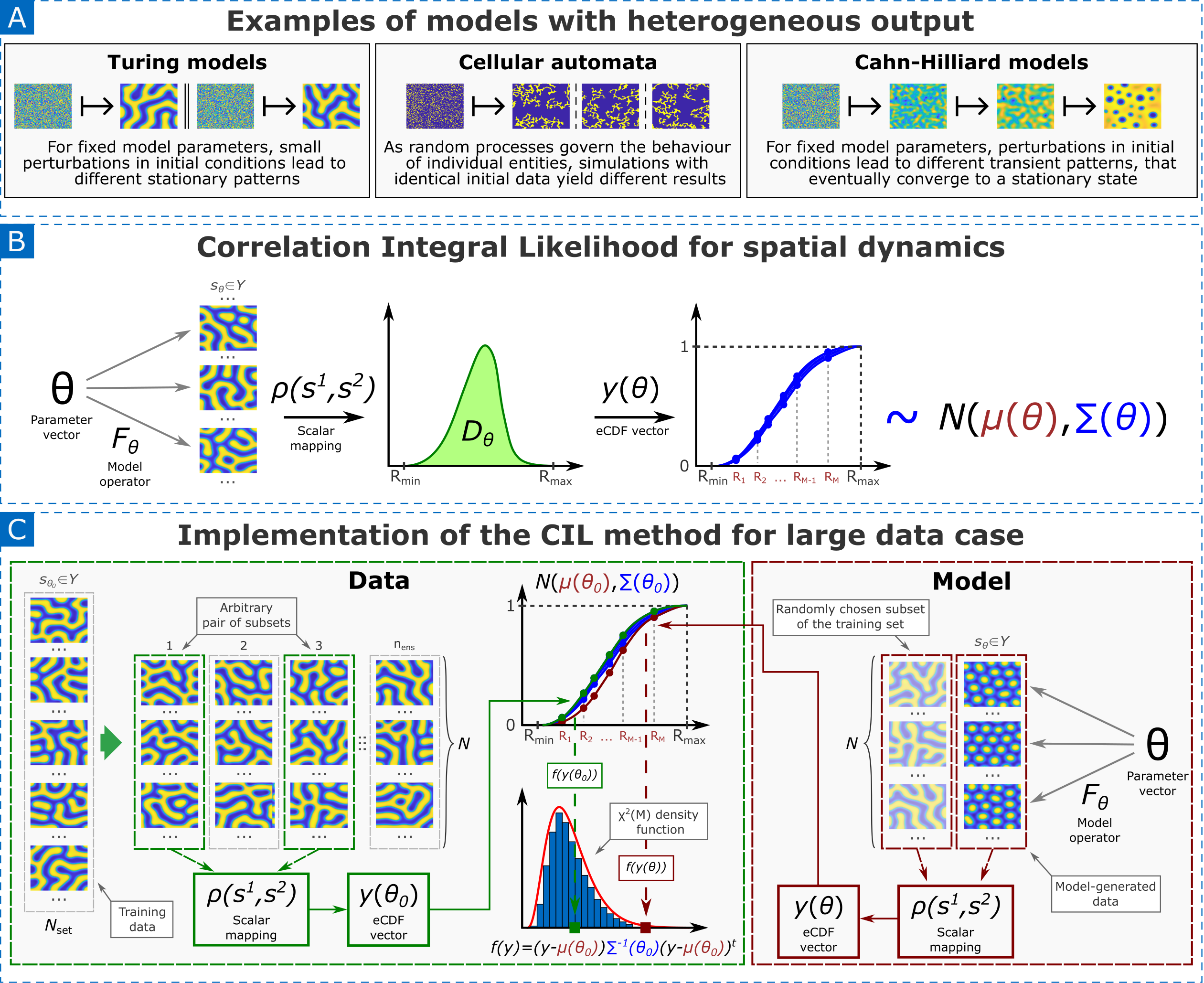
- Methodology Introduces the Correlation Integral Likelihood (CIL) framework, a unified approach for parameter estimation in systems with heterogeneous or chaotic dynamics (e.g., pattern formation, individual-based models), moving beyond classical Bayesian methods.
- Methodology Proposes integration of deterministic gradient flow methods within the CIL framework to enhance inference efficiency and accuracy, compared to traditional stochastic sampling (e.g., MCMC).
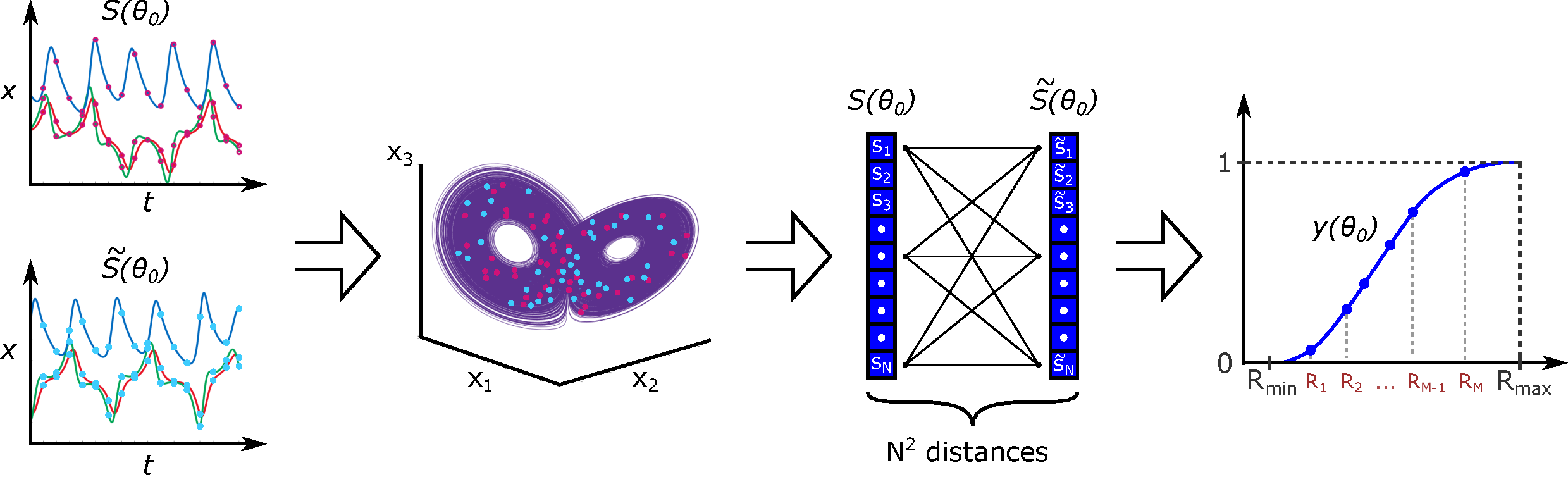
- Theory Generalizes the concept of correlation dimension from chaos theory to construct a robust metric for comparing the global geometric structure of model outputs (e.g., attractors, spatial patterns) rather than relying on unstable pointwise comparisons.
主要结论
- The CIL method provides a theoretically grounded framework for parameter estimation in systems where solution heterogeneity (e.g., in Turing patterns or chaotic attractors) makes conventional likelihoods ineffective.
- Integrating deterministic gradient flow sampling with the CIL framework can potentially enhance computational efficiency and inference accuracy compared to purely stochastic methods like MCMC, especially for high-dimensional parameter spaces.
- The approach enables reliable model calibration and validation even with incomplete, noisy, or single-snapshot data, advancing the predictive capability and mechanistic understanding of complex biological systems.
摘要: Calibrating mathematical models of biological processes is essential for achieving predictive accuracy and gaining mechanistic insight. However, this task remains challenging due to limited and noisy data, significant biological variability, and the computational complexity of the models themselves. In this method's article, we explore a range of approaches for parameter inference in partial differential equation (PDE) models of biological systems. We introduce a unified mathematical framework, the Correlation Integral Likelihood (CIL) method, for parameter estimation in systems exhibiting heterogeneous or chaotic dynamics, encompassing both pattern formation models and individual-based models. Departing from classical Bayesian inverse problem methodologies, we motivate the development of the CIL method, demonstrate its versatility, and highlight illustrative applications within mathematical biology. Furthermore, we compare stochastic sampling strategies, such as Markov Chain Monte Carlo (MCMC), with deterministic gradient flow approaches, highlighting how these methods can be integrated within the proposed framework to enhance inference performance. Our work provides a practical and theoretically grounded toolbox for researchers seeking to calibrate complex biological models using incomplete, noisy, or heterogeneous data, thereby advancing both the predictive capability and mechanistic understanding of such systems.