Paper List
-
The Effective Reproduction Number in the Kermack-McKendrick model with age of infection and reinfection
This paper addresses the challenge of accurately estimating the time-varying effective reproduction number ℛ(t) in epidemics by incorporating two crit...
-
Covering Relations in the Poset of Combinatorial Neural Codes
This work addresses the core challenge of navigating the complex poset structure of neural codes to systematically test the conjecture linking convex ...
-
Collective adsorption of pheromones at the water-air interface
This paper addresses the core challenge of understanding how amphiphilic pheromones, previously assumed to be transported in the gas phase, can be sta...
-
pHapCompass: Probabilistic Assembly and Uncertainty Quantification of Polyploid Haplotype Phase
This paper addresses the core challenge of accurately assembling polyploid haplotypes from sequencing data, where read assignment ambiguity and an exp...
-
Setting up for failure: automatic discovery of the neural mechanisms of cognitive errors
This paper addresses the core challenge of automating the discovery of biologically plausible recurrent neural network (RNN) dynamics that can replica...
-
Influence of Object Affordance on Action Language Understanding: Evidence from Dynamic Causal Modeling Analysis
This study addresses the core challenge of moving beyond correlational evidence to establish the *causal direction* and *temporal dynamics* of how obj...
-
Revealing stimulus-dependent dynamics through statistical complexity
This paper addresses the core challenge of detecting stimulus-specific patterns in neural population dynamics that remain hidden to traditional variab...
-
Exactly Solvable Population Model with Square-Root Growth Noise and Cell-Size Regulation
This paper addresses the fundamental gap in understanding how microscopic growth fluctuations, specifically those with size-dependent (square-root) no...
PanFoMa: A Lightweight Foundation Model and Benchmark for Pan-Cancer
Extracted from affiliations in the content snippet (specific institutions not fully listed in provided text)
30秒速读
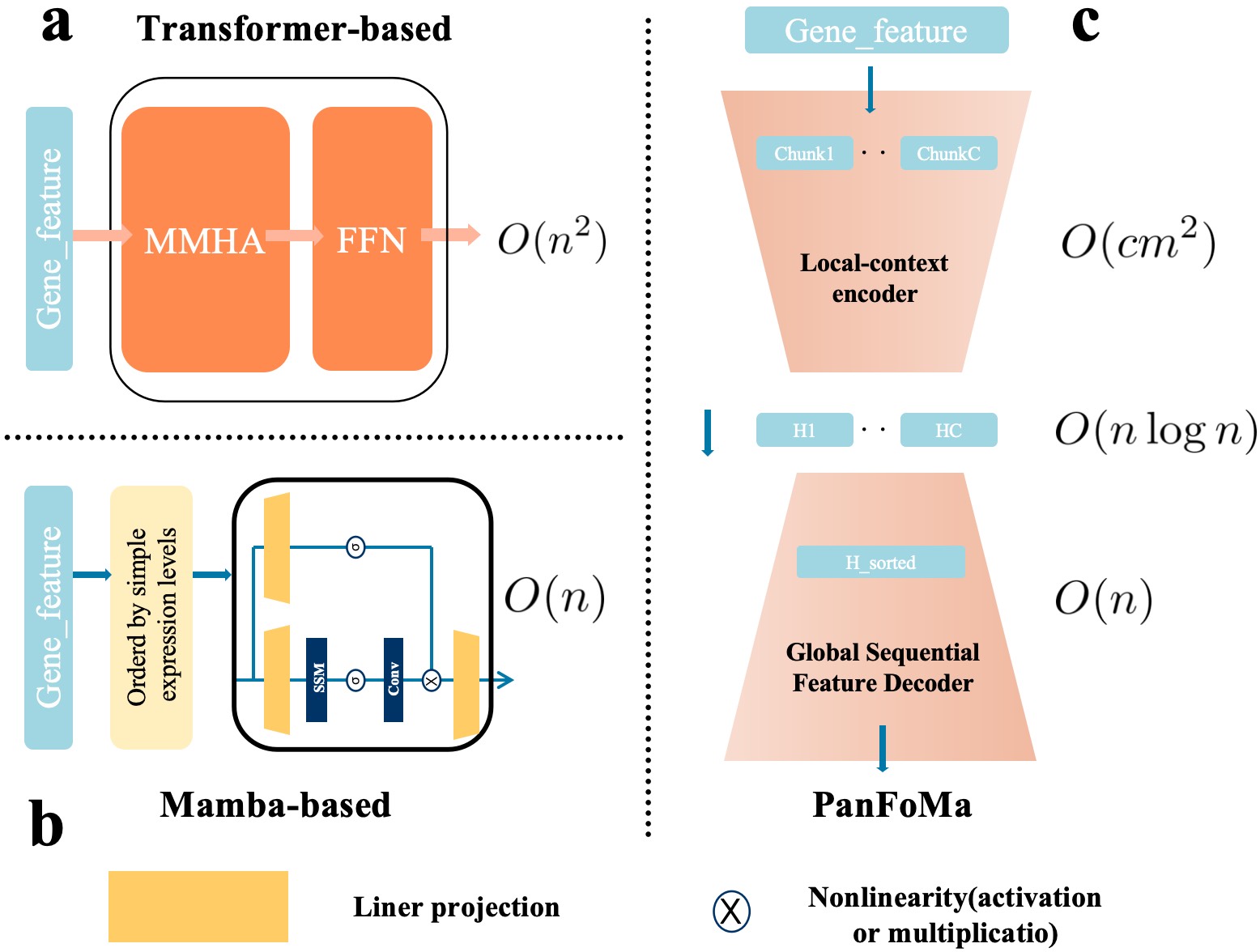
IN SHORT: This paper addresses the dual challenge of achieving computational efficiency without sacrificing accuracy in whole-transcriptome single-cell representation learning for pan-cancer analysis, moving beyond the limitations of pure Transformer or Mamba architectures.
核心创新
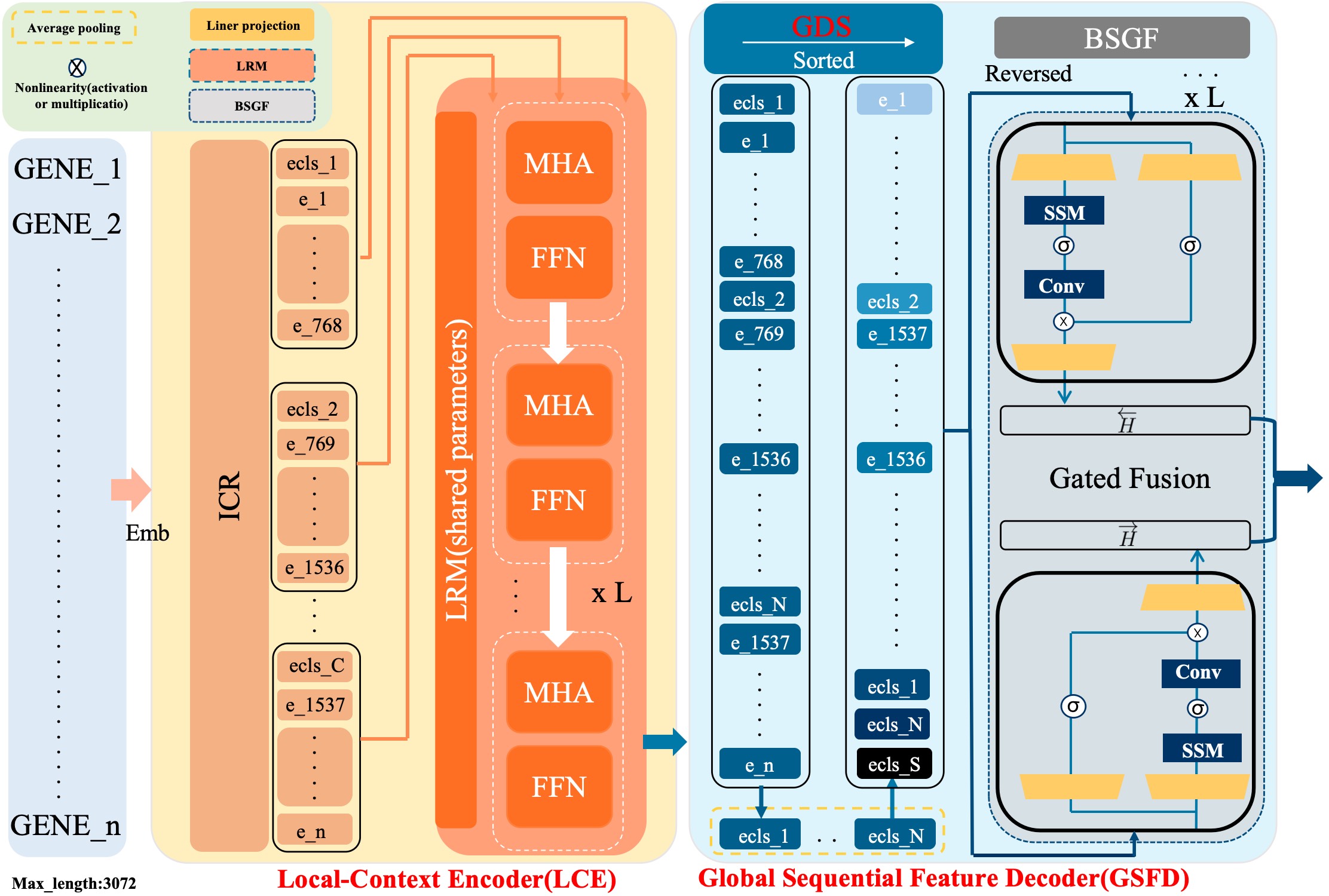
- Methodology Proposes a novel hybrid architecture (PanFoMa) that decouples local gene interaction modeling (via a lightweight, chunked Transformer encoder) from global context integration (via a bidirectional Mamba decoder), achieving O(C·M² + N log N) complexity.
- Methodology Introduces a Global-informed Dynamic Sorting (GDS) mechanism that adaptively orders genes for the Mamba decoder based on a learned global cell state vector, moving beyond static, heuristic gene ordering (e.g., by mean expression).
- Biology Constructs and releases PanFoMaBench, a large-scale, rigorously curated pan-cancer single-cell benchmark comprising over 3.5 million high-quality cells across 33 cancer subtypes from 23 tissues, addressing the lack of comprehensive evaluation resources.
主要结论
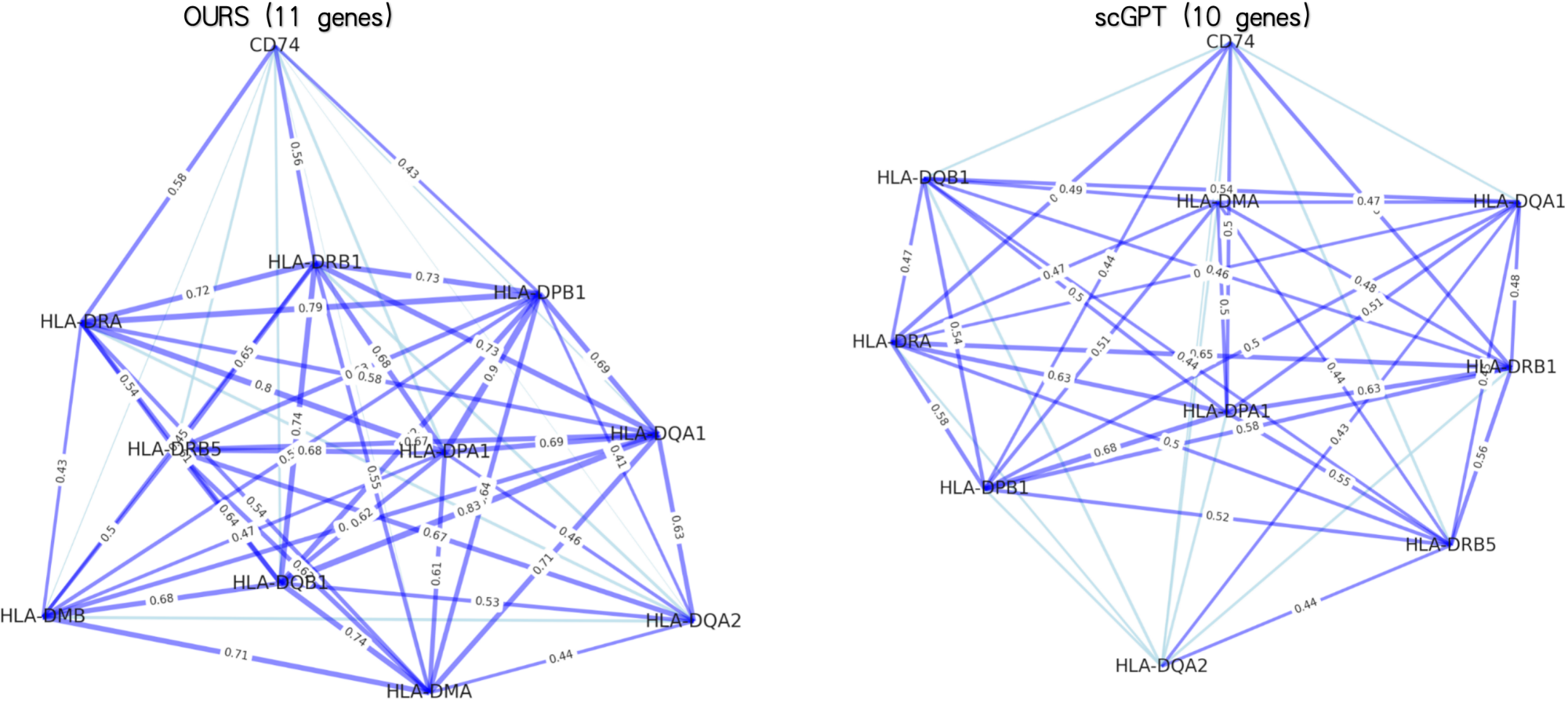
- PanFoMa achieves state-of-the-art pan-cancer classification accuracy of 94.74% (ACC) and 92.5% (Macro-F1) on PanFoMaBench, outperforming GeneFormer by +3.5% ACC and +4.0% F1.
- The model demonstrates superior generalizability across foundational tasks, showing improvements of +7.4% in cell type annotation, +4.0% in batch integration, and +3.1% in multi-omics integration over baselines.
- The hybrid local-global design and dynamic sorting are validated as effective, enabling efficient processing of full transcriptome-scale data (~3000 genes) while capturing both fine-grained local interactions and broad global regulatory patterns.
摘要: Single-cell RNA sequencing (scRNA-seq) is essential for decoding tumor heterogeneity. However, pan-cancer research still faces two key challenges: learning discriminative and efficient single-cell representations, and establishing a comprehensive evaluation benchmark. In this paper, we introduce PanFoMa, a lightweight hybrid neural network that combines the strengths of Transformers and state-space models to achieve a balance between performance and efficiency. PanFoMa consists of a front-end local-context encoder with shared self-attention layers to capture complex, order-independent gene interactions; and a back-end global sequential feature decoder that efficiently integrates global context using a linear-time state-space model. This modular design preserves the expressive power of Transformers while leveraging the scalability of Mamba to enable transcriptome modeling, effectively capturing both local and global regulatory signals. To enable robust evaluation, we also construct a large-scale pan-cancer single-cell benchmark, PanFoMaBench, containing over 3.5 million high-quality cells across 33 cancer subtypes, curated through a rigorous preprocessing pipeline. Experimental results show that PanFoMa outperforms state-of-the-art models on our pan-cancer benchmark (+4.0%) and across multiple public tasks, including cell type annotation (+7.4%), batch integration (+4.0%) and multi-omics integration (+3.1%). The code is available at https://github.com/Xiaoshui-Huang/PanFoMa.