Paper List
-
Discovery of a Hematopoietic Manifold in scGPT Yields a Method for Extracting Performant Algorithms from Biological Foundation Model Internals
This work addresses the core challenge of extracting reusable, interpretable, and high-performance biological algorithms from the opaque internal repr...
-
MS2MetGAN: Latent-space adversarial training for metabolite–spectrum matching in MS/MS database search
This paper addresses the critical bottleneck in metabolite identification: the generation of high-quality negative training samples that are structura...
-
Toward Robust, Reproducible, and Widely Accessible Intracranial Language Brain-Computer Interfaces: A Comprehensive Review of Neural Mechanisms, Hardware, Algorithms, Evaluation, Clinical Pathways and Future Directions
This review addresses the core challenge of fragmented and heterogeneous evidence that hinders the clinical translation of intracranial language BCIs,...
-
Less Is More in Chemotherapy of Breast Cancer
通过纳入细胞周期时滞和竞争项,解决了现有肿瘤-免疫模型的过度简化问题,以定量比较化疗方案。
-
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
This paper addresses the critical bottleneck of GPU memory limitations that restrict AlphaFold 3-like models to processing only a few thousand residue...
-
Open Biomedical Knowledge Graphs at Scale: Construction, Federation, and AI Agent Access with Samyama Graph Database
This paper addresses the core pain point of fragmented biomedical data by constructing and federating large-scale, open knowledge graphs to enable sea...
-
Predictive Analytics for Foot Ulcers Using Time-Series Temperature and Pressure Data
This paper addresses the critical need for continuous, real-time monitoring of diabetic foot health by developing an unsupervised anomaly detection fr...
-
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, wh...
Approximate Bayesian Inference on Mechanisms of Network Growth and Evolution
Harvard T.H. Chan School of Public Health
30秒速读
IN SHORT: This paper addresses the core challenge of inferring the relative contributions of multiple, simultaneous generative mechanisms in network formation when the true likelihood is intractable.
核心创新
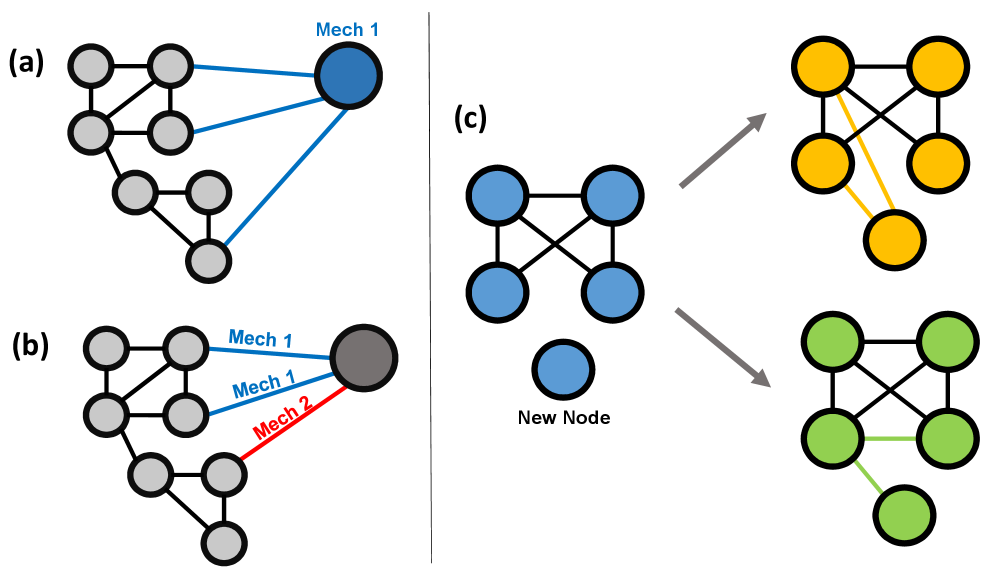
- Methodology Proposes an event-wise mixture-of-mechanisms model that assigns generative rules (e.g., Preferential Attachment, Random Attachment) to each edge formation event, rather than to nodes, increasing model flexibility and realism.
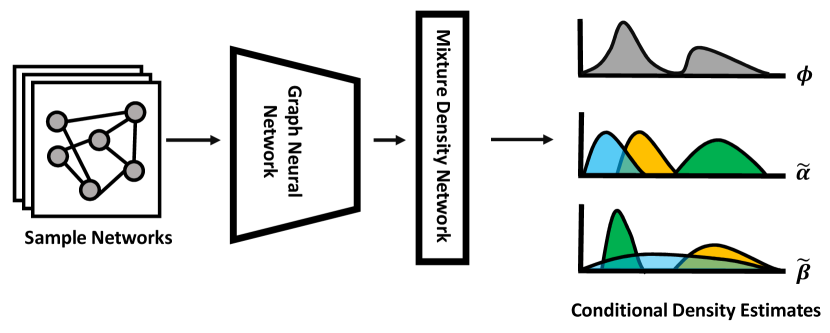
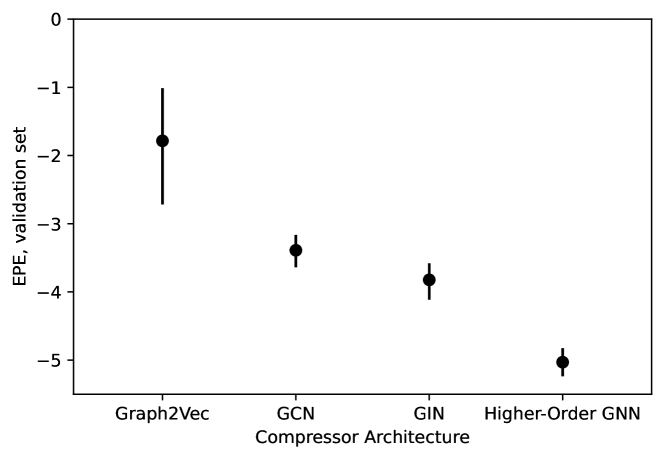
- Methodology Introduces a novel GNN-MDN (Graph Neural Network - Mixture Density Network) architecture that automatically learns informative, low-dimensional network embeddings for conditional density estimation, bypassing the need for manually specified summary statistics.
- Theory Formalizes a unified framework that incorporates both growth mechanisms (adding nodes/edges) and evolution mechanisms (modifying existing edges), allowing the model to capture a wider range of network dynamics like triangle formation.
主要结论
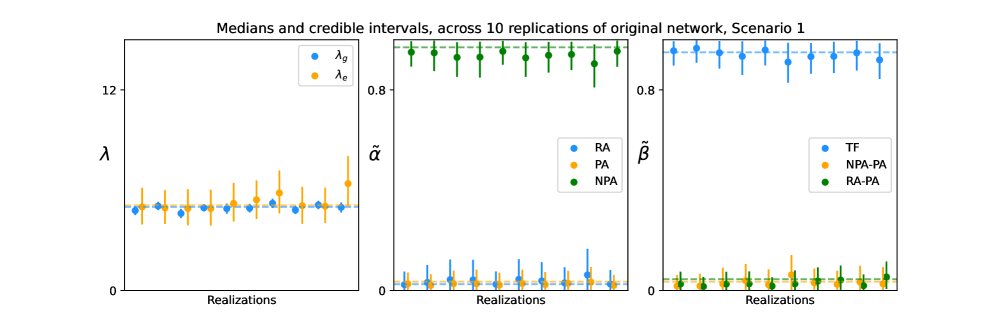
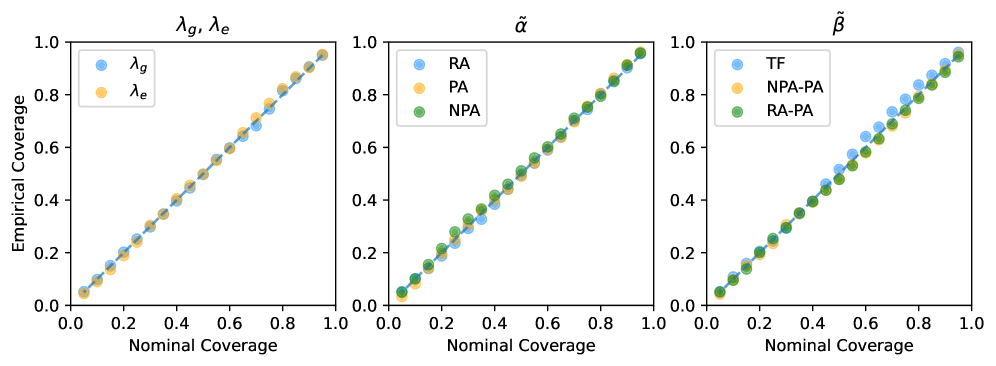
- The proposed GNN-MDN method provides valid approximate Bayesian inference, demonstrated via simulation studies showing that the 95% credible intervals achieve nominal coverage (e.g., containing the true parameter values).
- The event-wise model successfully infers dominant mechanisms in simulated scenarios; for instance, it accurately recovers a weight vector of (0.95, 0.025, 0.025) for a scenario where Preferential Attachment is the primary growth mechanism.
- The method is applicable to real-world networks, providing interpretable decompositions of their formation processes into quantifiable contributions from mechanisms like Random Attachment, Preferential Attachment, and Triangle Formation.
摘要: Mechanistic models can provide an intuitive and interpretable explanation of network growth by specifying a set of generative rules. These rules can be defined by domain knowledge about real-world mechanisms governing network growth or may be designed to facilitate the appearance of certain network motifs. In the formation of real-world networks, multiple mechanisms may be simultaneously involved; it is then important to understand the relative contribution of each of these mechanisms. In this paper, we propose the use of a conditional density estimator, augmented with a graph neural network, to perform inference on a flexible mixture of network-forming mechanisms. This event-wise mixture-of-mechanisms model assigns mechanisms to each edge formation event rather than stipulating node-level mechanisms, thus allowing for an explanation of the network generation process, as well as the dynamic evolution of the network over time. We demonstrate that our approximate Bayesian approach yields valid inferences for the relative weights of the mechanisms in our model, and we utilize this method to investigate the mechanisms behind the formation of a variety of real-world networks.