Paper List
-
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcomi...
-
Incorporating indel channels into average-case analysis of seed-chain-extend
This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigoro...
-
Competition, stability, and functionality in excitatory-inhibitory neural circuits
This paper addresses the core challenge of extending interpretable energy-based frameworks to biologically realistic asymmetric neural networks, where...
-
Enhancing Clinical Note Generation with ICD-10, Clinical Ontology Knowledge Graphs, and Chain-of-Thought Prompting Using GPT-4
This paper addresses the core challenge of generating accurate and clinically relevant patient notes from sparse inputs (ICD codes and basic demograph...
-
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited his...
-
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring ...
-
Generating a Contact Matrix for Aged Care Settings in Australia: an agent-based model study
This study addresses the critical gap in understanding heterogeneous contact patterns within aged care facilities, where existing population-level con...
-
Emergent Spatiotemporal Dynamics in Large-Scale Brain Networks with Next Generation Neural Mass Models
This work addresses the core challenge of understanding how complex, brain-wide spatiotemporal patterns emerge from the interaction of biophysically d...
Approximate Bayesian Inference on Mechanisms of Network Growth and Evolution
Harvard T.H. Chan School of Public Health
30秒速读
IN SHORT: This paper addresses the core challenge of inferring the relative contributions of multiple, simultaneous generative mechanisms in network formation when the true likelihood is intractable.
核心创新
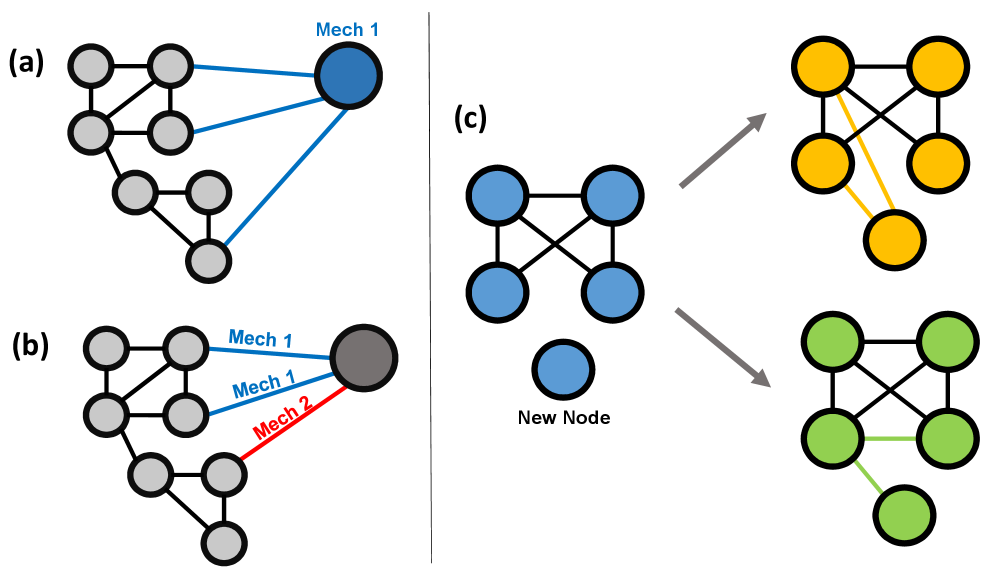
- Methodology Proposes an event-wise mixture-of-mechanisms model that assigns generative rules (e.g., Preferential Attachment, Random Attachment) to each edge formation event, rather than to nodes, increasing model flexibility and realism.
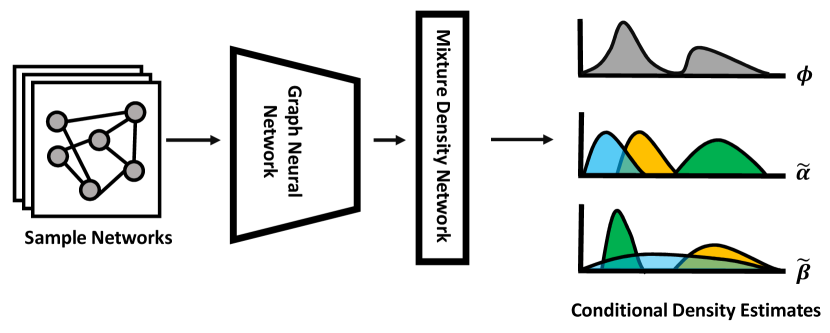
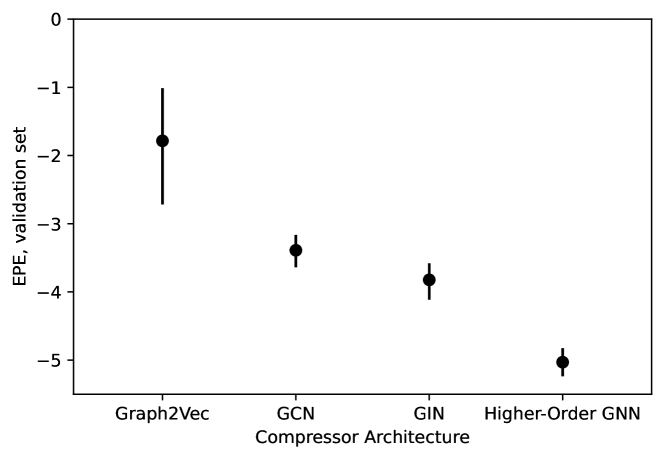
- Methodology Introduces a novel GNN-MDN (Graph Neural Network - Mixture Density Network) architecture that automatically learns informative, low-dimensional network embeddings for conditional density estimation, bypassing the need for manually specified summary statistics.
- Theory Formalizes a unified framework that incorporates both growth mechanisms (adding nodes/edges) and evolution mechanisms (modifying existing edges), allowing the model to capture a wider range of network dynamics like triangle formation.
主要结论
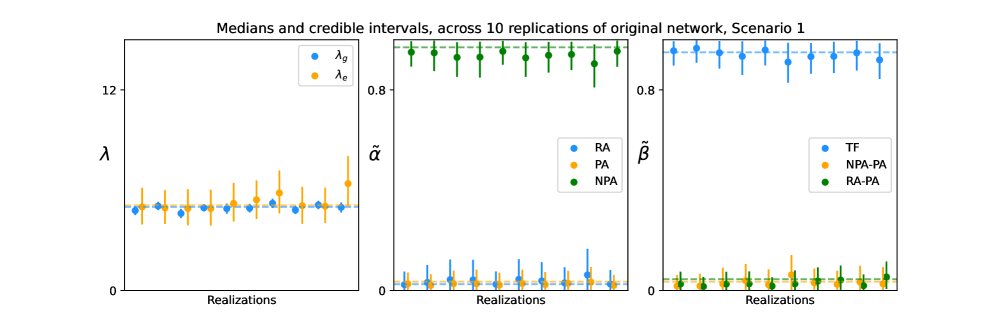
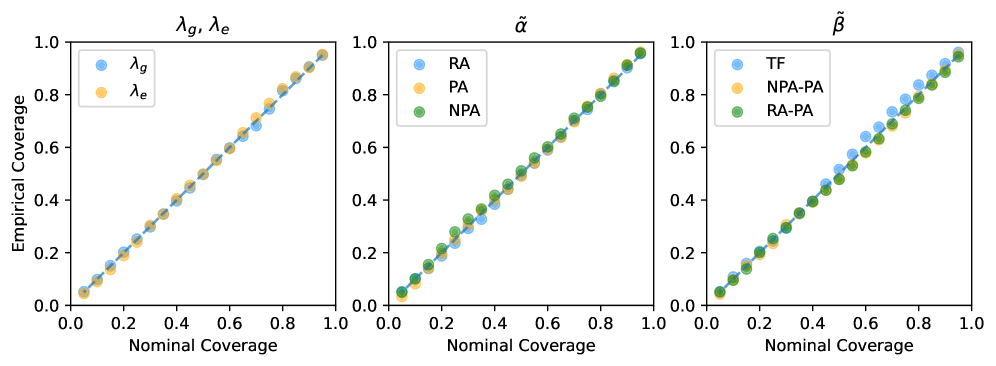
- The proposed GNN-MDN method provides valid approximate Bayesian inference, demonstrated via simulation studies showing that the 95% credible intervals achieve nominal coverage (e.g., containing the true parameter values).
- The event-wise model successfully infers dominant mechanisms in simulated scenarios; for instance, it accurately recovers a weight vector of (0.95, 0.025, 0.025) for a scenario where Preferential Attachment is the primary growth mechanism.
- The method is applicable to real-world networks, providing interpretable decompositions of their formation processes into quantifiable contributions from mechanisms like Random Attachment, Preferential Attachment, and Triangle Formation.
摘要: Mechanistic models can provide an intuitive and interpretable explanation of network growth by specifying a set of generative rules. These rules can be defined by domain knowledge about real-world mechanisms governing network growth or may be designed to facilitate the appearance of certain network motifs. In the formation of real-world networks, multiple mechanisms may be simultaneously involved; it is then important to understand the relative contribution of each of these mechanisms. In this paper, we propose the use of a conditional density estimator, augmented with a graph neural network, to perform inference on a flexible mixture of network-forming mechanisms. This event-wise mixture-of-mechanisms model assigns mechanisms to each edge formation event rather than stipulating node-level mechanisms, thus allowing for an explanation of the network generation process, as well as the dynamic evolution of the network over time. We demonstrate that our approximate Bayesian approach yields valid inferences for the relative weights of the mechanisms in our model, and we utilize this method to investigate the mechanisms behind the formation of a variety of real-world networks.