Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
Modulation of DNA rheology by a transcription factor that forms aging microgels
University of Edinburgh | University of Glasgow | MRC Human Genetics Unit | WPI-SKCM2, Hiroshima University
30秒速读
IN SHORT: This work addresses the fundamental question of how the transcription factor NANOG, essential for embryonic stem cell pluripotency, physically regulates gene expression beyond simple DNA binding, by revealing its ability to form self-limiting, aging microgels that modulate DNA rheology.
核心创新
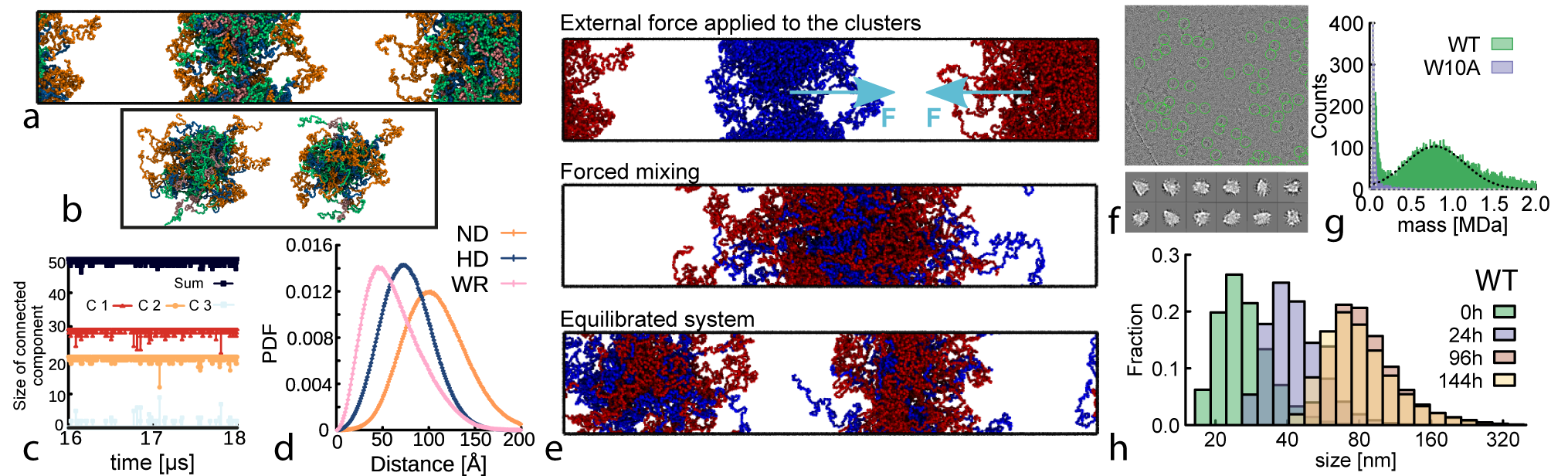
- Methodology First demonstration that a transcription factor (NANOG) forms self-limiting micelle-like clusters (~22-25 monomers) with exposed DNA-binding domains, acting as transient cross-linkers for DNA molecules.
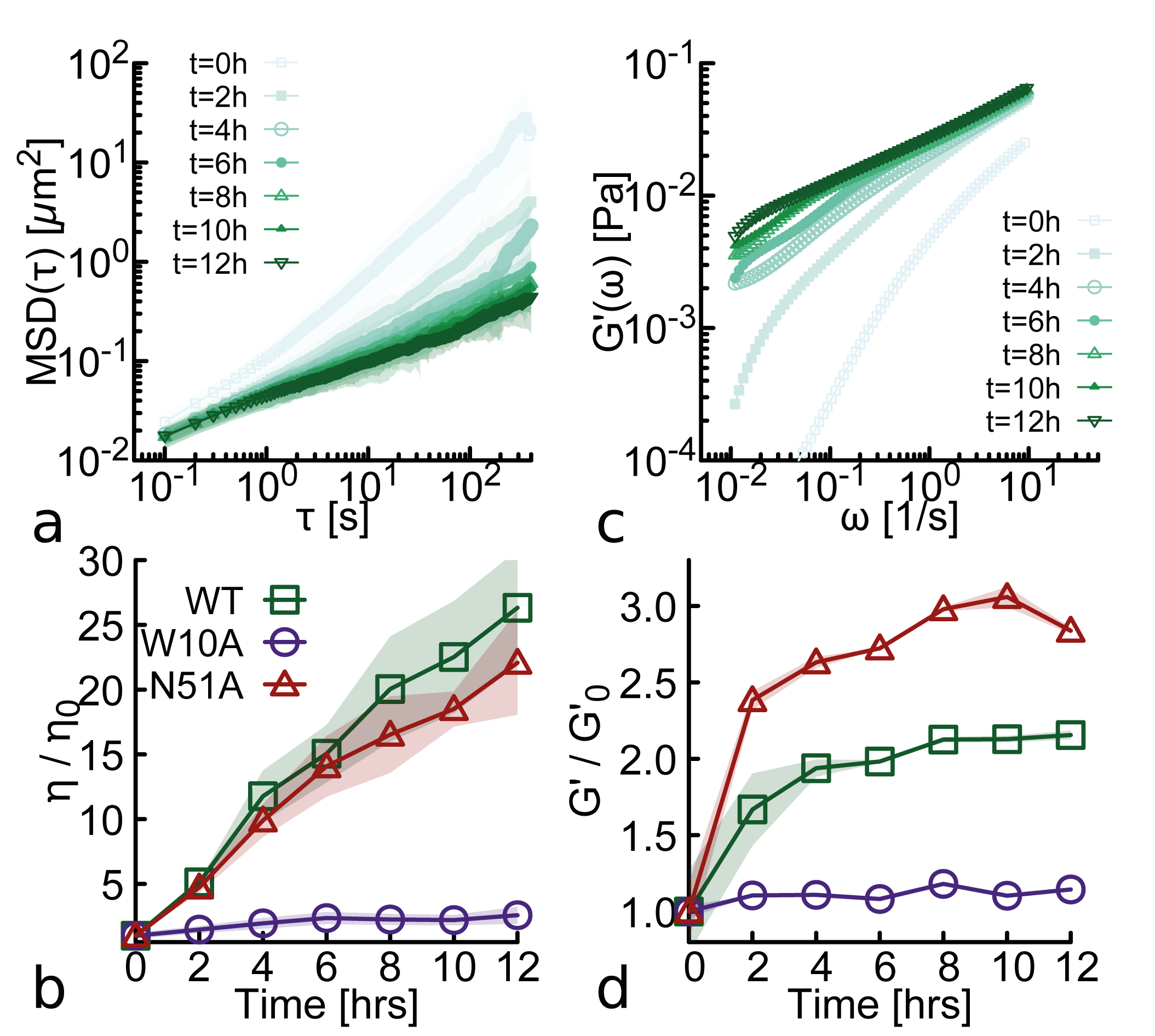
- Biology Discovery of an aging microgel formation by NANOG, where viscoelasticity increases over time (10,000-fold viscosity increase over 12h), driven by its intrinsically disordered tryptophan-rich (WR) domain.
- Theory Proposes a novel 'rheological gene regulation' paradigm: NANOG may regulate gene expression not by large-scale chromatin reorganization, but by stabilizing and restricting the *dynamics* of key regulatory sites via aging condensates, potentially ingraining mechanical memory.
主要结论
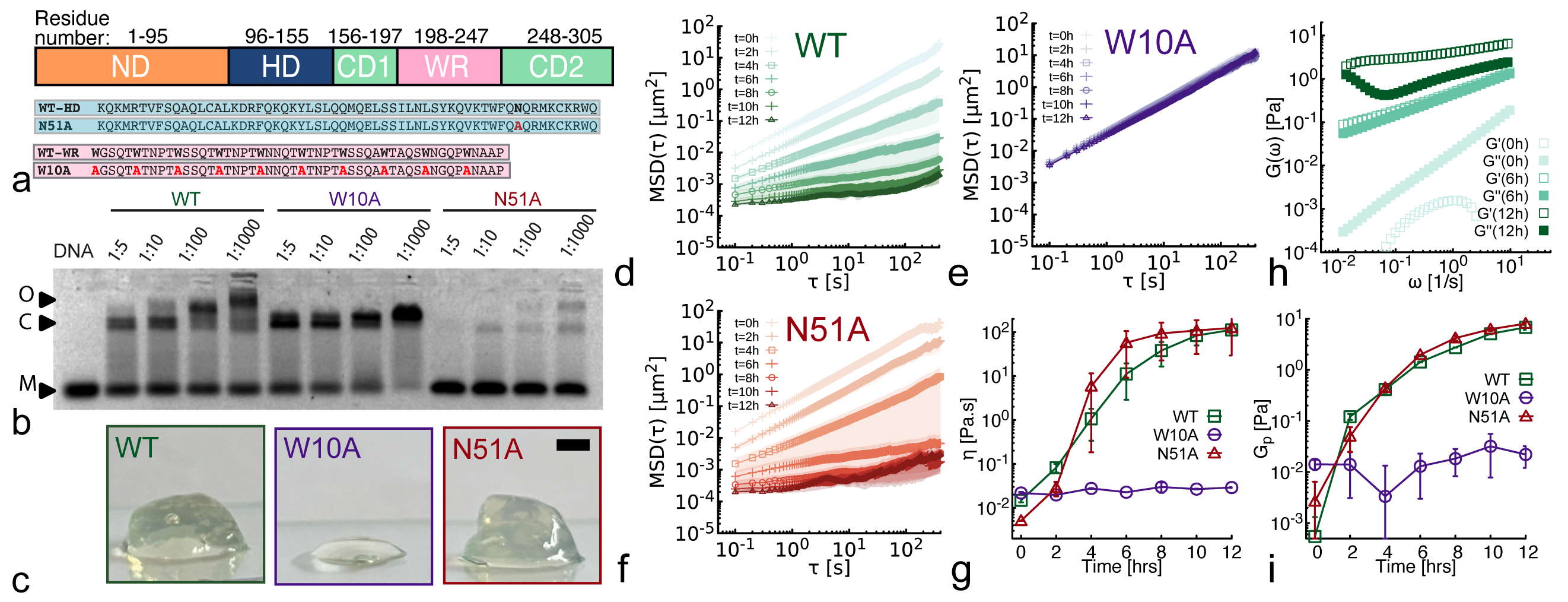
- Wild-type NANOG forms macroscopic aging gels (10,000-fold viscosity increase over 12h at 37°C) and self-limiting micelle-like clusters (~22-25 proteins), while the oligomerization-deficient mutant (W10A) does not.
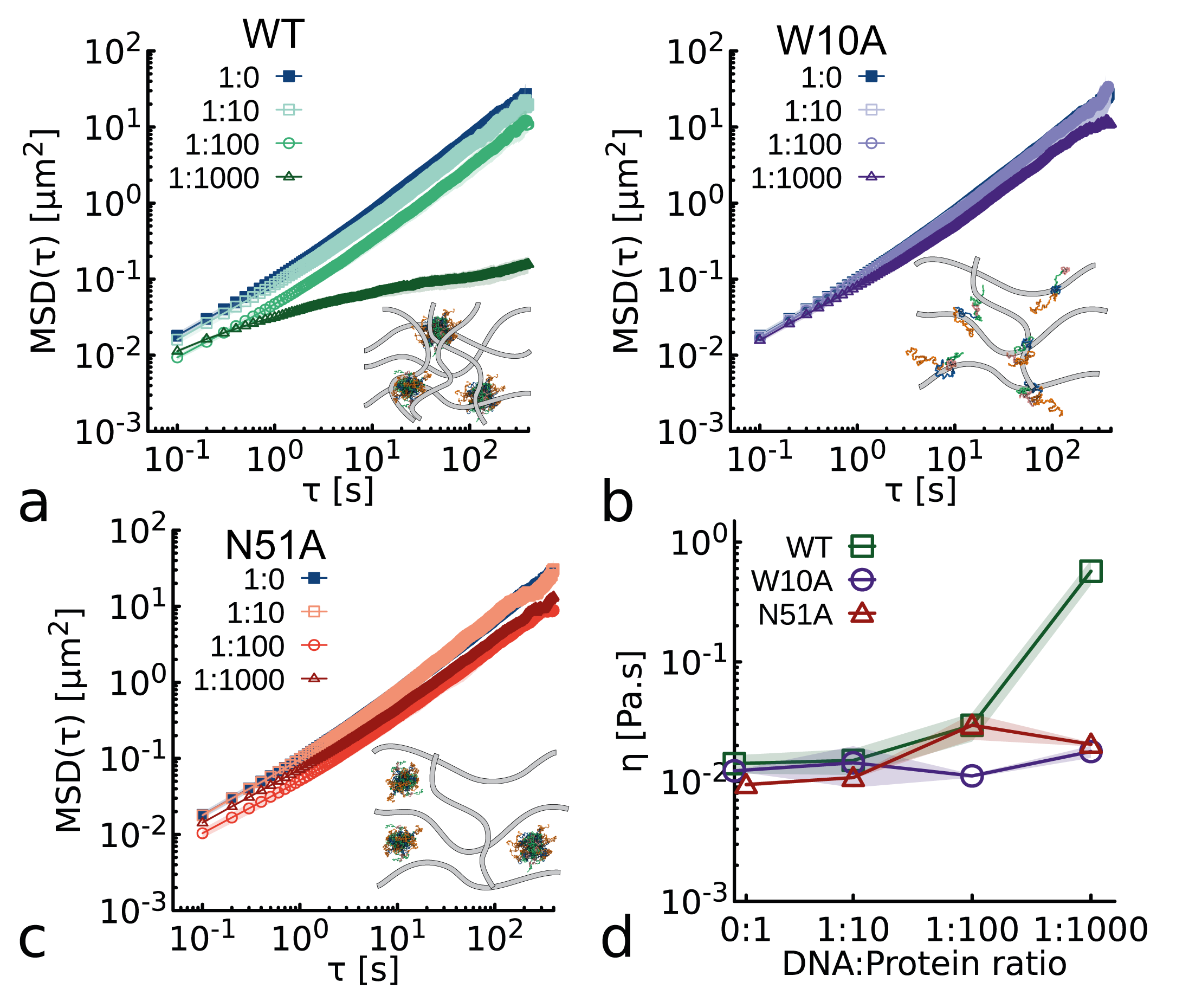
- Both clustering (via WR domain) and DNA binding (via homeodomain) are required for NANOG to act as an effective DNA cross-linker, significantly enhancing the viscoelasticity of entangled DNA solutions (observed in WT but not in W10A or DNA-binding-deficient N51A mutants).
- Aging (increasing viscoelasticity over time) occurs in NANOG-DNA solutions for both WT and the DNA-binding-deficient N51A mutant, indicating that oligomerization alone is sufficient to drive this slow restructuring toward gel-like states.
摘要: Proteins and nucleic acids form non-Newtonian liquids with complex rheological properties that contribute to their function in vivo. Here we investigate the rheology of the transcription factor NANOG, a key protein in sustaining embryonic stem cell self-renewal. We discover that at high concentrations NANOG forms macroscopic aging gels through its intrinsically disordered tryptophan-rich domain. By combining molecular dynamics simulations, mass photometry and Cryo-EM, we also discover that NANOG forms self-limiting micelle-like clusters which expose their DNA-binding domains. In dense solutions of DNA, NANOG micelle-like structures stabilize inter-molecular entanglements and crosslinks, forming microgel-like structures. Our findings suggest that NANOG may contribute to regulate gene expression in a unconventional way: by restricting and stabilizing genome dynamics at key transcriptional sites through the formation of an aging microgel-like structure, potentially enabling mechanical memory in the gene network.