Paper List
-
Evolutionarily Stable Stackelberg Equilibrium
通过要求追随者策略对突变入侵具有鲁棒性,弥合了斯塔克尔伯格领导力模型与演化稳定性之间的鸿沟。
-
Recovering Sparse Neural Connectivity from Partial Measurements: A Covariance-Based Approach with Granger-Causality Refinement
通过跨多个实验会话累积协方差统计,实现从部分记录到完整神经连接性的重建。
-
Atomic Trajectory Modeling with State Space Models for Biomolecular Dynamics
ATMOS通过提供一个基于SSM的高效框架,用于生物分子的原子级轨迹生成,弥合了计算昂贵的MD模拟与时间受限的深度生成模型之间的差距。
-
Slow evolution towards generalism in a model of variable dietary range
通过证明是种群统计噪声(而非确定性动力学)驱动了模式形成和泛化食性的演化,解决了间接竞争下物种形成的悖论。
-
Grounded Multimodal Retrieval-Augmented Drafting of Radiology Impressions Using Case-Based Similarity Search
通过将印象草稿基于检索到的历史病例,并采用明确引用和基于置信度的拒绝机制,解决放射学报告生成中的幻觉问题。
-
Unified Policy–Value Decomposition for Rapid Adaptation
通过双线性分解在策略和价值函数之间共享低维目标嵌入,实现对新颖任务的零样本适应。
-
Mathematical Modeling of Cancer–Bacterial Therapy: Analysis and Numerical Simulation via Physics-Informed Neural Networks
提供了一个严格的、无网格的PINN框架,用于模拟和分析细菌癌症疗法中复杂的、空间异质的相互作用。
-
Sample-Efficient Adaptation of Drug-Response Models to Patient Tumors under Strong Biological Domain Shift
通过从无标记分子谱中学习可迁移表征,利用最少的临床数据实现患者药物反应的有效预测。
scCluBench: Comprehensive Benchmarking of Clustering Algorithms for Single-Cell RNA Sequencing
Not specified in provided content
30秒速读
IN SHORT: This paper addresses the critical gap of fragmented and non-standardized benchmarking in single-cell RNA-seq clustering, which hinders objective comparison and selection of appropriate methods for specific biological contexts.
核心创新
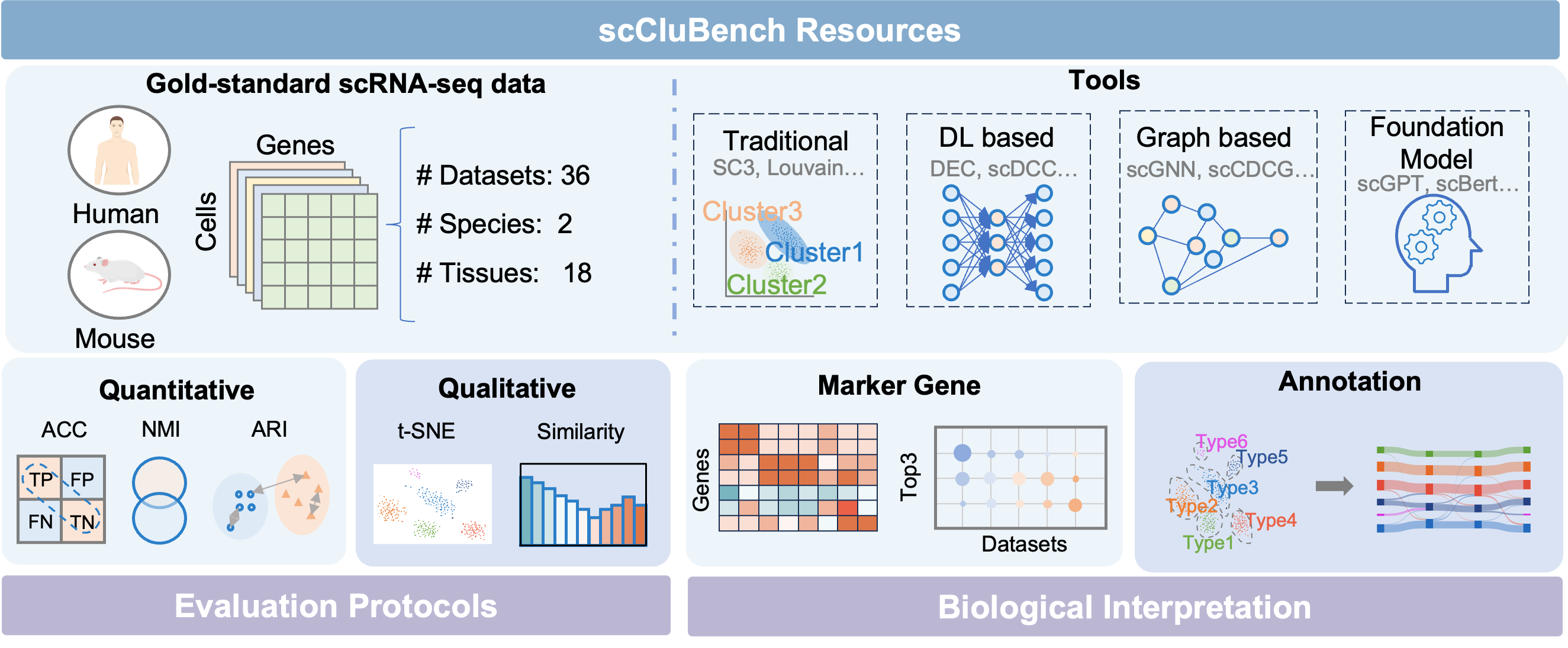
- Methodology Introduces scCluBench, the first comprehensive benchmarking framework that systematically evaluates 16 clustering methods across four categories (traditional, deep learning-based, graph-based, and foundation models) on 36 standardized datasets.
- Methodology Establishes standardized protocols for biological interpretation, including reproducible pipelines for marker gene identification and two distinct cell type annotation approaches (best-mapping and marker-overlap), validated with gold-standard references.
- Methodology Provides a unified and modular benchmarking workflow covering data preprocessing, clustering, and annotation with standardized input-output formats, ensuring reproducibility and fair comparison.
主要结论
- scCDCG (a cut-informed graph embedding model) achieved the highest average clustering accuracy (81.29 ± 1.45) across 36 datasets, outperforming other graph-based, deep learning, and traditional methods.
- Biological foundation models (scGPT, GeneFormer, GeneCompass) showed strong performance in classification tasks (e.g., scGPT achieved 98.14% ACC on Sapiens Ear Crista Ampullaris) but underperformed in direct clustering, highlighting a trade-off between general representation and task-specific optimization.
- The benchmark reveals method-specific limitations: traditional methods struggle with sparse data, deep learning models may fail to capture cell relationships, and graph-based models can suffer from over-smoothing, while most methods decouple embedding learning from clustering optimization.
摘要: Cell clustering is crucial for uncovering cellular heterogeneity in single-cell RNA sequencing (scRNA-seq) data by identifying cell types and marker genes. Despite its importance, benchmarks for scRNA-seq clustering methods remain fragmented, often lacking standardized protocols and failing to incorporate recent advances in artificial intelligence. To fill these gaps, we present scCluBench, a comprehensive benchmark of clustering algorithms for scRNA-seq data. First, scCluBench provides 36 scRNA-seq datasets collected from diverse public sources, covering multiple tissues, which are uniformly processed and standardized to ensure consistency for systematic evaluation and downstream analyses. To evaluate performance, we collect and reproduce a range of scRNA-seq clustering methods, including traditional, deep learning-based, graph-based, and biological foundation models. We comprehensively evaluate each method both quantitatively and qualitatively, using core performance metrics as well as visualization analyses. Furthermore, we construct representative downstream biological tasks, such as marker gene identification and cell type annotation, to further assess the practical utility. scCluBench then investigates the performance differences and applicability boundaries of various clustering models across diverse analytical tasks, systematically assessing their robustness and scalability in real-world scenarios. Overall, scCluBench offers a standardized and user-friendly benchmark for scRNA-seq clustering, with curated datasets, unified evaluation protocols, and transparent analyses, facilitating informed method selection and providing valuable insights into model generalizability and application scope.222All datasets, code, and the Extended version for scCluBench are available at the link: https://github.com/XPgogogo/scCluBench. More details for each stage are provided in the extended version.