Paper List
-
Discovery of a Hematopoietic Manifold in scGPT Yields a Method for Extracting Performant Algorithms from Biological Foundation Model Internals
This work addresses the core challenge of extracting reusable, interpretable, and high-performance biological algorithms from the opaque internal repr...
-
MS2MetGAN: Latent-space adversarial training for metabolite–spectrum matching in MS/MS database search
This paper addresses the critical bottleneck in metabolite identification: the generation of high-quality negative training samples that are structura...
-
Toward Robust, Reproducible, and Widely Accessible Intracranial Language Brain-Computer Interfaces: A Comprehensive Review of Neural Mechanisms, Hardware, Algorithms, Evaluation, Clinical Pathways and Future Directions
This review addresses the core challenge of fragmented and heterogeneous evidence that hinders the clinical translation of intracranial language BCIs,...
-
Less Is More in Chemotherapy of Breast Cancer
通过纳入细胞周期时滞和竞争项,解决了现有肿瘤-免疫模型的过度简化问题,以定量比较化疗方案。
-
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
This paper addresses the critical bottleneck of GPU memory limitations that restrict AlphaFold 3-like models to processing only a few thousand residue...
-
Open Biomedical Knowledge Graphs at Scale: Construction, Federation, and AI Agent Access with Samyama Graph Database
This paper addresses the core pain point of fragmented biomedical data by constructing and federating large-scale, open knowledge graphs to enable sea...
-
Predictive Analytics for Foot Ulcers Using Time-Series Temperature and Pressure Data
This paper addresses the critical need for continuous, real-time monitoring of diabetic foot health by developing an unsupervised anomaly detection fr...
-
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, wh...
Few-shot Protein Fitness Prediction via In-context Learning and Test-time Training
Department of Systems Biology, Harvard Medical School | Department of Biology, University of Copenhagen | Machine Intelligence, Novo Nordisk A/S | Microsoft Research, Cambridge, MA, USA | Dept. of Applied Mathematics and Computer Science, Technical University of Denmark
30秒速读
IN SHORT: This paper addresses the core challenge of accurately predicting protein fitness with only a handful of experimental observations, where data collection is prohibitively expensive and label availability is severely limited.
核心创新
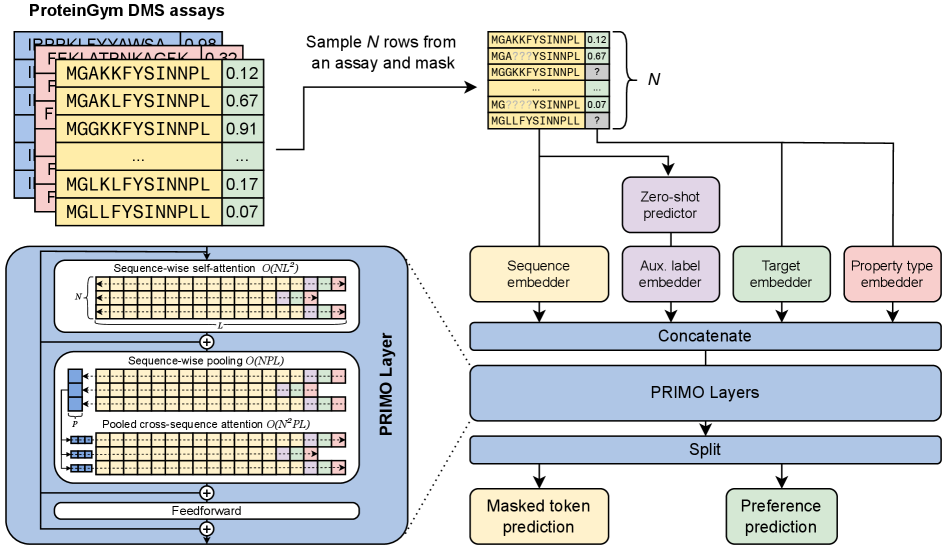
- Methodology Introduces PRIMO, a novel transformer-based framework that uniquely combines in-context learning with test-time training for few-shot protein fitness prediction.
- Methodology Proposes a hybrid masked token reconstruction objective with a preference-based loss function, enabling effective learning from sparse experimental labels across diverse assays.
- Methodology Develops a lightweight pooling attention mechanism that handles both substitution and indel mutations while maintaining computational efficiency, overcoming limitations of previous methods.
主要结论
- PRIMO with test-time training (TTT) achieves state-of-the-art few-shot performance, improving from a zero-shot Spearman correlation of 0.51 to 0.67 with 128 shots, outperforming Gaussian Process (0.56) and Ridge Regression (0.63) baselines.
- The framework demonstrates broad applicability across protein properties including stability (0.77 correlation with TTT), enzymatic activity (0.61), fluorescence (0.30), and binding (0.69), handling both substitution and indel mutations.
- PRIMO's performance highlights the critical importance of proper data splitting to avoid inflated results, as demonstrated by the 0.4 correlation inflation on RL40A_YEAST when using Metalic's overlapping train-test split.
摘要: Accurately predicting protein fitness with minimal experimental data is a persistent challenge in protein engineering. We introduce PRIMO (PRotein In-context Mutation Oracle), a transformer-based framework that leverages in-context learning and test-time training to adapt rapidly to new proteins and assays without large task-specific datasets. By encoding sequence information, auxiliary zero-shot predictions, and sparse experimental labels from many assays as a unified token set in a pre-training masked-language modeling paradigm, PRIMO learns to prioritize promising variants through a preference-based loss function. Across diverse protein families and properties—including both substitution and indel mutations—PRIMO outperforms zero-shot and fully supervised baselines. This work underscores the power of combining large-scale pre-training with efficient test-time adaptation to tackle challenging protein design tasks where data collection is expensive and label availability is limited.