Paper List
-
Formation of Artificial Neural Assemblies by Biologically Plausible Inhibition Mechanisms
This work addresses the core limitation of the Assembly Calculus model—its fixed-size, biologically implausible k-WTA selection process—by introducing...
-
How to make the most of your masked language model for protein engineering
This paper addresses the critical bottleneck of efficiently sampling high-quality, diverse protein sequences from Masked Language Models (MLMs) for pr...
-
Module control in youth symptom networks across COVID-19
This paper addresses the core challenge of distinguishing whether a prolonged societal stressor (COVID-19) fundamentally reorganizes the architecture ...
-
JEDI: Jointly Embedded Inference of Neural Dynamics
This paper addresses the core challenge of inferring context-dependent neural dynamics from noisy, high-dimensional recordings using a single unified ...
-
ATP Level and Phosphorylation Free Energy Regulate Trigger-Wave Speed and Critical Nucleus Size in Cellular Biochemical Systems
This work addresses the core challenge of quantitatively predicting how the cellular energy state (ATP level and phosphorylation free energy) governs ...
-
Packaging Jupyter notebooks as installable desktop apps using LabConstrictor
This paper addresses the core pain point of ensuring Jupyter notebook reproducibility and accessibility across different computing environments, parti...
-
SNPgen: Phenotype-Supervised Genotype Representation and Synthetic Data Generation via Latent Diffusion
This paper addresses the core challenge of generating privacy-preserving synthetic genotype data that maintains both statistical fidelity and downstre...
-
Continuous Diffusion Transformers for Designing Synthetic Regulatory Elements
This paper addresses the challenge of efficiently generating novel, cell-type-specific regulatory DNA sequences with high predicted activity while min...
Few-shot Protein Fitness Prediction via In-context Learning and Test-time Training
Department of Systems Biology, Harvard Medical School | Department of Biology, University of Copenhagen | Machine Intelligence, Novo Nordisk A/S | Microsoft Research, Cambridge, MA, USA | Dept. of Applied Mathematics and Computer Science, Technical University of Denmark
30秒速读
IN SHORT: This paper addresses the core challenge of accurately predicting protein fitness with only a handful of experimental observations, where data collection is prohibitively expensive and label availability is severely limited.
核心创新
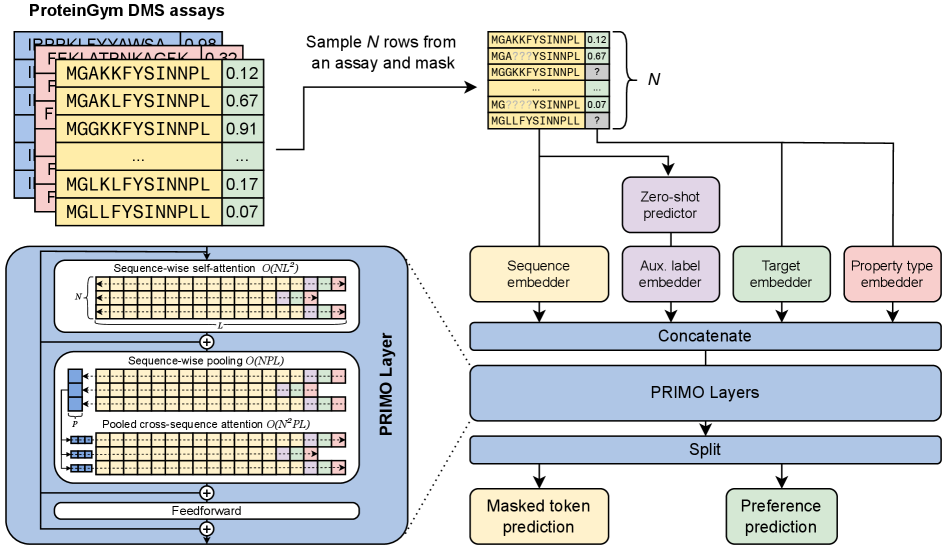
- Methodology Introduces PRIMO, a novel transformer-based framework that uniquely combines in-context learning with test-time training for few-shot protein fitness prediction.
- Methodology Proposes a hybrid masked token reconstruction objective with a preference-based loss function, enabling effective learning from sparse experimental labels across diverse assays.
- Methodology Develops a lightweight pooling attention mechanism that handles both substitution and indel mutations while maintaining computational efficiency, overcoming limitations of previous methods.
主要结论
- PRIMO with test-time training (TTT) achieves state-of-the-art few-shot performance, improving from a zero-shot Spearman correlation of 0.51 to 0.67 with 128 shots, outperforming Gaussian Process (0.56) and Ridge Regression (0.63) baselines.
- The framework demonstrates broad applicability across protein properties including stability (0.77 correlation with TTT), enzymatic activity (0.61), fluorescence (0.30), and binding (0.69), handling both substitution and indel mutations.
- PRIMO's performance highlights the critical importance of proper data splitting to avoid inflated results, as demonstrated by the 0.4 correlation inflation on RL40A_YEAST when using Metalic's overlapping train-test split.
摘要: Accurately predicting protein fitness with minimal experimental data is a persistent challenge in protein engineering. We introduce PRIMO (PRotein In-context Mutation Oracle), a transformer-based framework that leverages in-context learning and test-time training to adapt rapidly to new proteins and assays without large task-specific datasets. By encoding sequence information, auxiliary zero-shot predictions, and sparse experimental labels from many assays as a unified token set in a pre-training masked-language modeling paradigm, PRIMO learns to prioritize promising variants through a preference-based loss function. Across diverse protein families and properties—including both substitution and indel mutations—PRIMO outperforms zero-shot and fully supervised baselines. This work underscores the power of combining large-scale pre-training with efficient test-time adaptation to tackle challenging protein design tasks where data collection is expensive and label availability is limited.