Paper List
-
Discovery of a Hematopoietic Manifold in scGPT Yields a Method for Extracting Performant Algorithms from Biological Foundation Model Internals
This work addresses the core challenge of extracting reusable, interpretable, and high-performance biological algorithms from the opaque internal repr...
-
MS2MetGAN: Latent-space adversarial training for metabolite–spectrum matching in MS/MS database search
This paper addresses the critical bottleneck in metabolite identification: the generation of high-quality negative training samples that are structura...
-
Toward Robust, Reproducible, and Widely Accessible Intracranial Language Brain-Computer Interfaces: A Comprehensive Review of Neural Mechanisms, Hardware, Algorithms, Evaluation, Clinical Pathways and Future Directions
This review addresses the core challenge of fragmented and heterogeneous evidence that hinders the clinical translation of intracranial language BCIs,...
-
Less Is More in Chemotherapy of Breast Cancer
通过纳入细胞周期时滞和竞争项,解决了现有肿瘤-免疫模型的过度简化问题,以定量比较化疗方案。
-
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
This paper addresses the critical bottleneck of GPU memory limitations that restrict AlphaFold 3-like models to processing only a few thousand residue...
-
Open Biomedical Knowledge Graphs at Scale: Construction, Federation, and AI Agent Access with Samyama Graph Database
This paper addresses the core pain point of fragmented biomedical data by constructing and federating large-scale, open knowledge graphs to enable sea...
-
Predictive Analytics for Foot Ulcers Using Time-Series Temperature and Pressure Data
This paper addresses the critical need for continuous, real-time monitoring of diabetic foot health by developing an unsupervised anomaly detection fr...
-
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, wh...
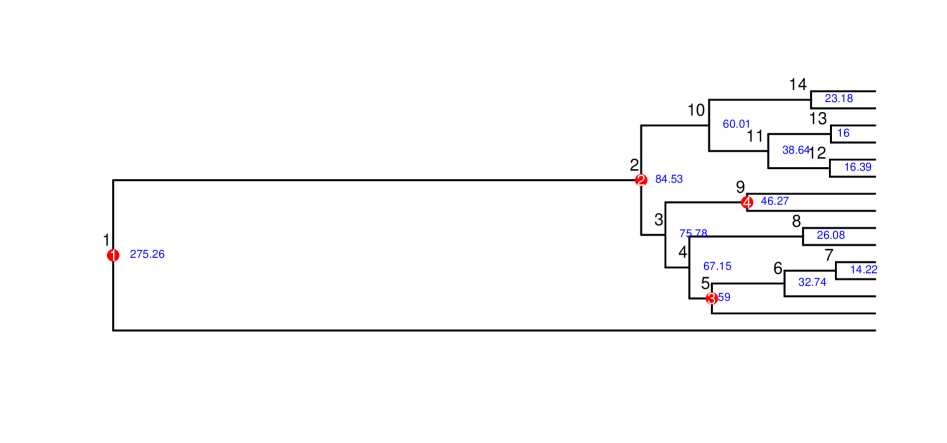
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
Department of Statistics, University of Georgia, Athens, 30601, USA
30秒速读
IN SHORT: This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimation for large phylogenomic datasets.
核心创新
- Methodology Introduces two Adjusted Pairwise Likelihood (APW) formulations (APW1 and APW2) that use asymptotic moment-matching weights to correct composite likelihoods within a Bayesian MCMC framework.
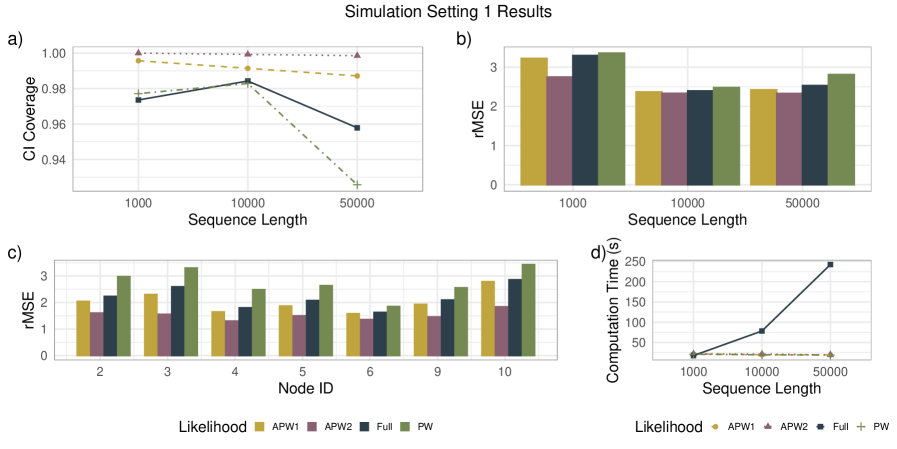
- Methodology Demonstrates that APW methods reduce computational cost by more than an order of magnitude compared to full-likelihood methods while maintaining comparable accuracy in node-age estimation.
- Methodology Shows that APW methods exhibit greater robustness to fossil misplacement and prior misspecification due to the reduced sensitivity of composite likelihoods to local calibration errors.
主要结论
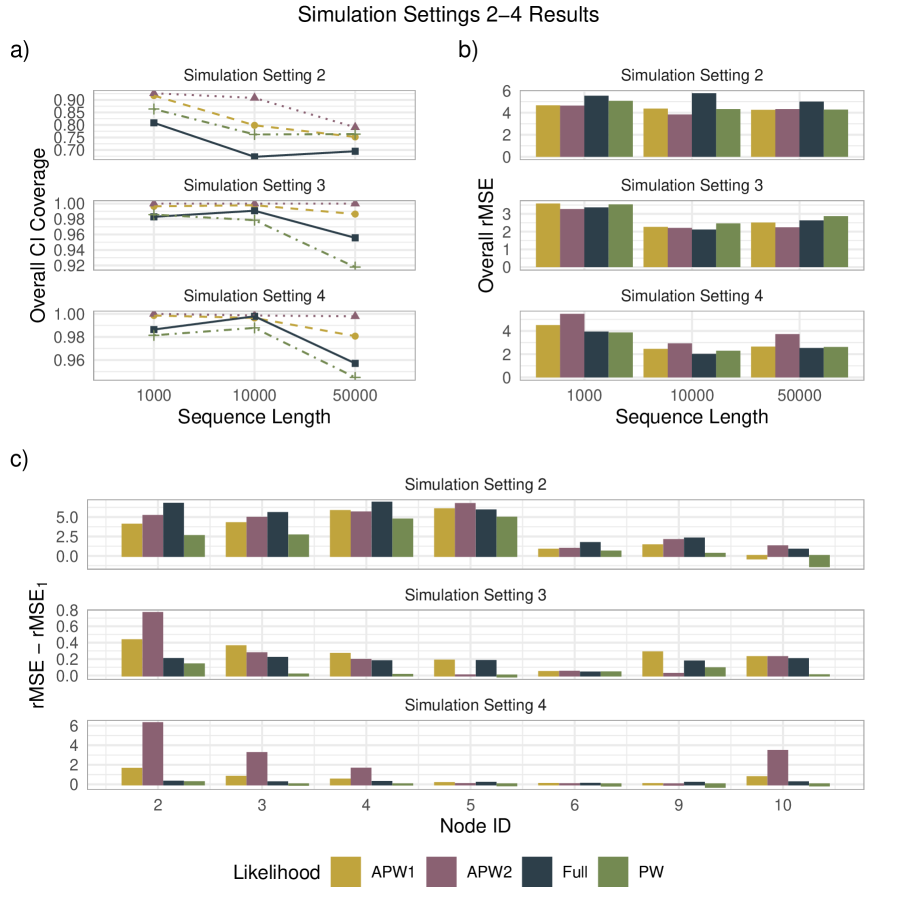
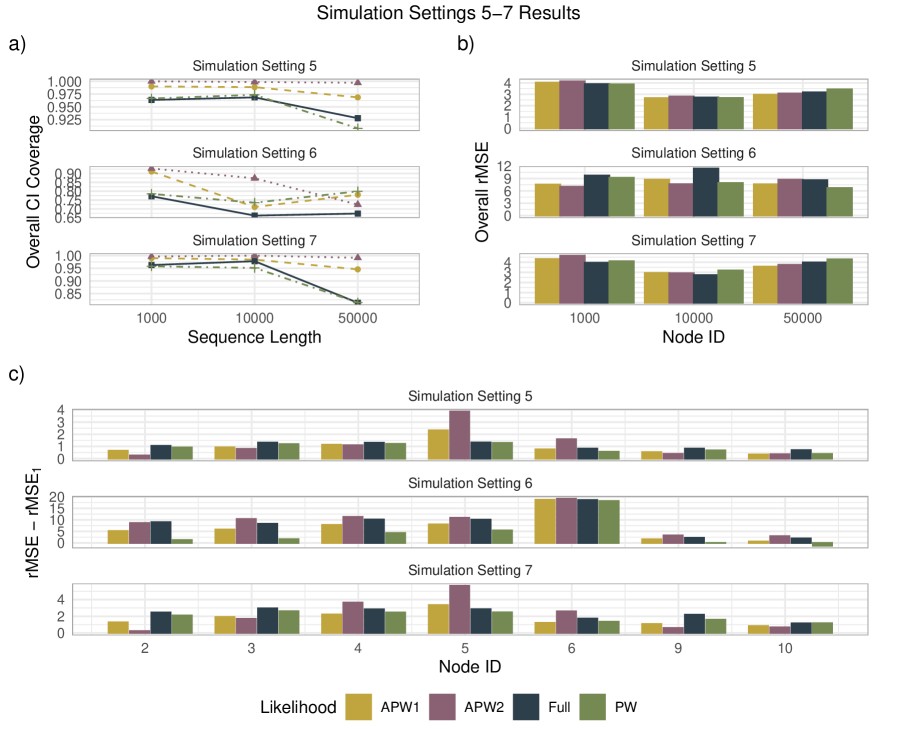
- APW methods produce node-age estimates statistically comparable to full-likelihood methods across diverse simulation scenarios, with reduced sensitivity to local calibration errors.
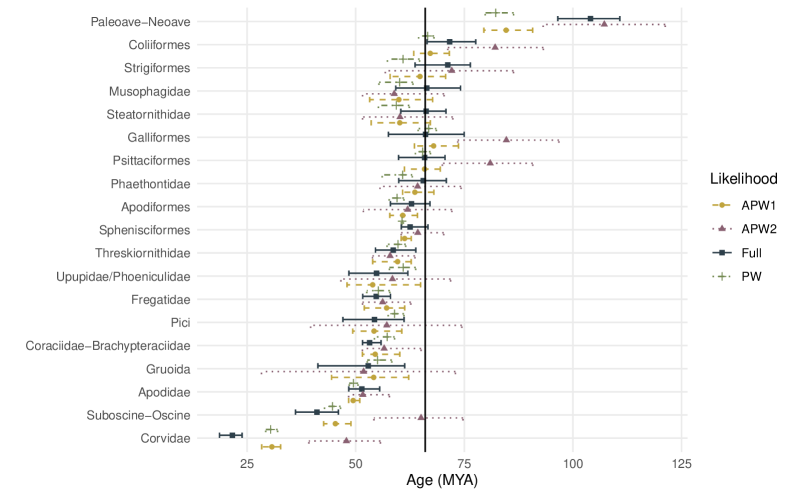
- Applied to a genome-scale avian dataset, APW recovered divergence time patterns consistent with recent studies while achieving a >10x reduction in computational cost.
- The robustness of APW to fossil misplacement stems from the composite likelihood's inherent property of being less sensitive to errors in individual calibration points, as demonstrated in simulations modeling various prior misspecifications.
摘要: Estimating divergence times from molecular sequence data is central to reconstructing the evolutionary history of lineages. Although Bayesian relaxed-clock methods provide a principled framework for incorporating fossil information, their dependence on repeated evaluations of the full phylogenetic likelihood makes them computationally demanding for large genomic datasets. Furthermore, because disagreements in divergence-time estimates often arise from uncertainty or error in fossil placement and prior specification, there is a need for methods that are both computationally efficient and robust to fossil-calibration uncertainty. In this study, we introduce fast and accurate alternatives based on the phylogenetic pairwise composite likelihood, presenting two adjusted pairwise likelihood (APW) formulations that employ asymptotic moment-matching weights to better approximate the behavior of the full likelihood within a Bayesian MCMC framework. Extensive simulations across diverse fossil-calibration scenarios show that APW methods produce node-age estimates comparable to those obtained from the full likelihood while offering greater robustness to fossil misplacement and prior misspecification, due to the reduced sensitivity of composite likelihoods to local calibration errors. Applied to a genome-scale dataset of modern birds, APW methods recover divergence time patterns consistent with recent studies, while reducing computational cost by more than an order of magnitude. Overall, our results demonstrate that adjusted pairwise likelihoods provide a calibration-robust and computationally efficient framework for Bayesian node dating, especially suited for large phylogenomic datasets and analyses in which fossil priors may be uncertain or imperfectly placed.