Paper List
-
Developing the PsyCogMetrics™ AI Lab to Evaluate Large Language Models and Advance Cognitive Science
This paper addresses the critical gap between sophisticated LLM evaluation needs and the lack of accessible, scientifically rigorous platforms that in...
-
Equivalence of approximation by networks of single- and multi-spike neurons
This paper resolves the fundamental question of whether single-spike spiking neural networks (SNNs) are inherently less expressive than multi-spike SN...
-
The neuroscience of transformers
提出了Transformer架构与皮层柱微环路之间的新颖计算映射,连接了现代AI与神经科学。
-
Framing local structural identifiability and observability in terms of parameter-state symmetries
This paper addresses the core challenge of systematically determining which parameters and states in a mechanistic ODE model can be uniquely inferred ...
-
Leveraging Phytolith Research using Artificial Intelligence
This paper addresses the critical bottleneck in phytolith research by automating the labor-intensive manual microscopy process through a multimodal AI...
-
Neural network-based encoding in free-viewing fMRI with gaze-aware models
This paper addresses the core challenge of building computationally efficient and ecologically valid brain encoding models for naturalistic vision by ...
-
Scalable DNA Ternary Full Adder Enabled by a Competitive Blocking Circuit
This paper addresses the core bottleneck of carry information attenuation and limited computational scale in DNA binary adders by introducing a scalab...
-
ELISA: An Interpretable Hybrid Generative AI Agent for Expression-Grounded Discovery in Single-Cell Genomics
This paper addresses the critical bottleneck of translating high-dimensional single-cell transcriptomic data into interpretable biological hypotheses ...
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
Department of Statistics, University of Georgia, Athens, 30601, USA
30秒速读
IN SHORT: This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimation for large phylogenomic datasets.
核心创新
- Methodology Introduces two Adjusted Pairwise Likelihood (APW) formulations (APW1 and APW2) that use asymptotic moment-matching weights to correct composite likelihoods within a Bayesian MCMC framework.
- Methodology Demonstrates that APW methods reduce computational cost by more than an order of magnitude compared to full-likelihood methods while maintaining comparable accuracy in node-age estimation.
- Methodology Shows that APW methods exhibit greater robustness to fossil misplacement and prior misspecification due to the reduced sensitivity of composite likelihoods to local calibration errors.
主要结论
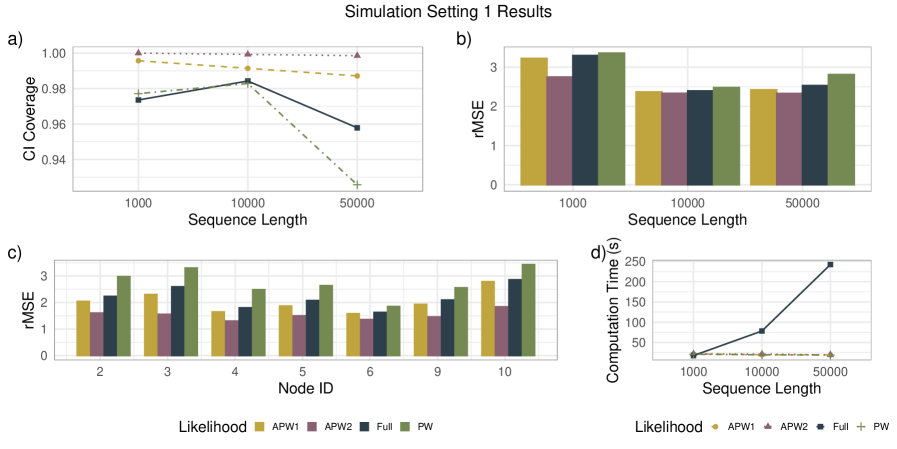
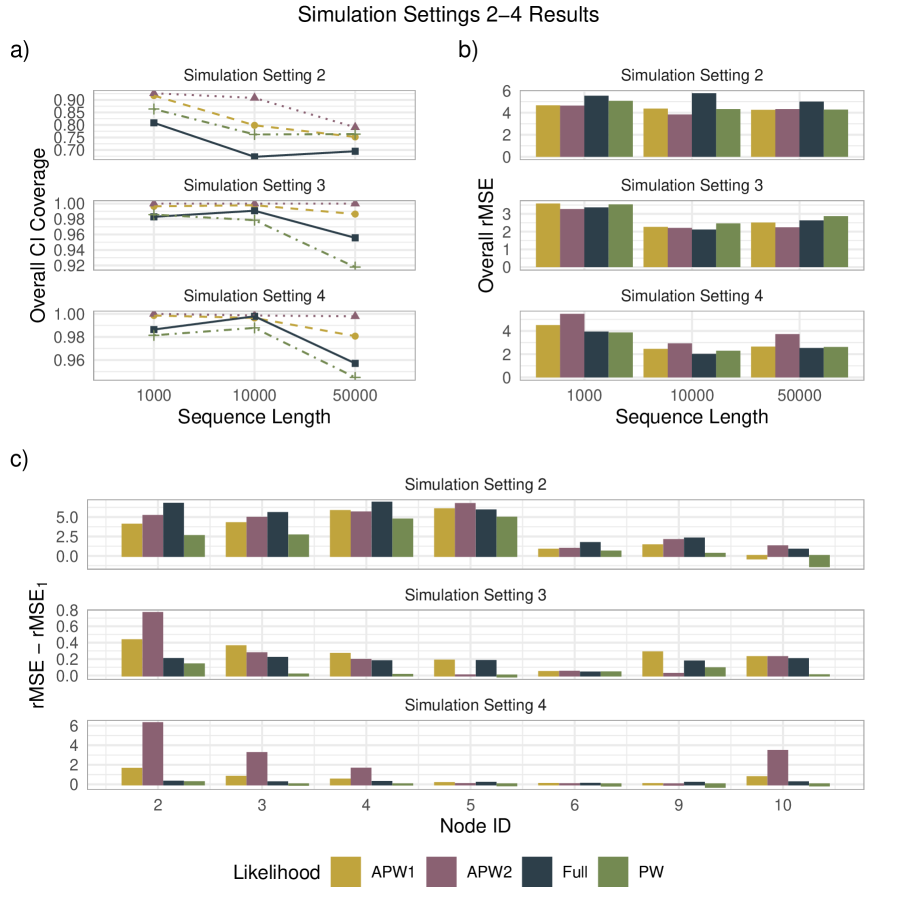
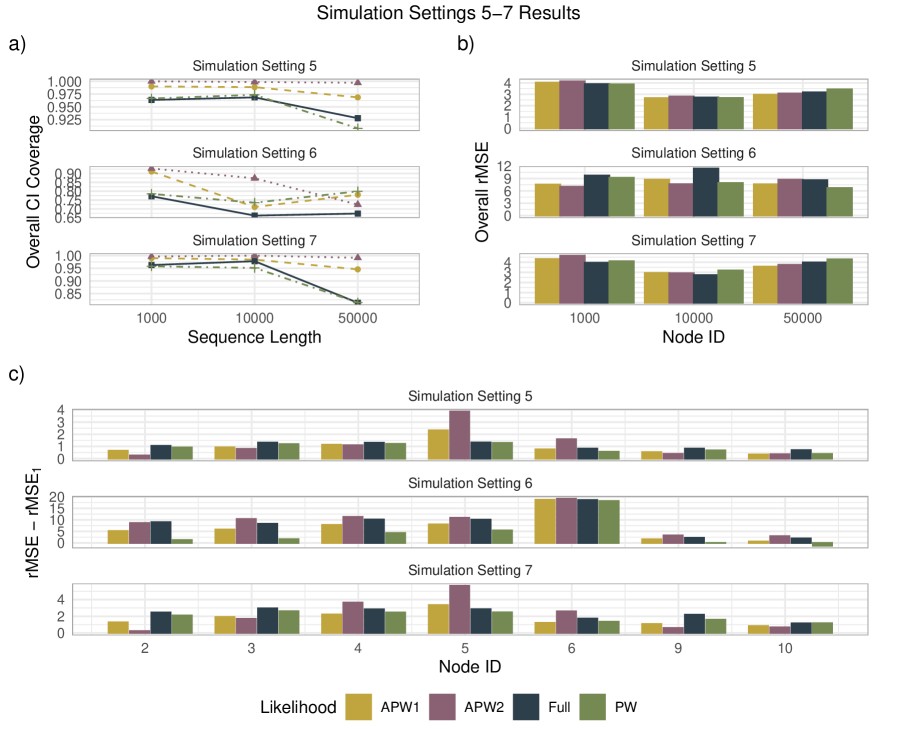
- APW methods produce node-age estimates statistically comparable to full-likelihood methods across diverse simulation scenarios, with reduced sensitivity to local calibration errors.
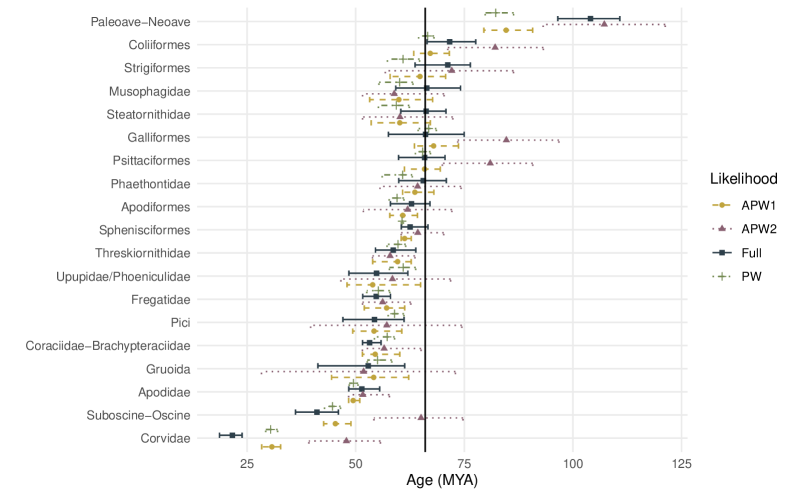
- Applied to a genome-scale avian dataset, APW recovered divergence time patterns consistent with recent studies while achieving a >10x reduction in computational cost.
- The robustness of APW to fossil misplacement stems from the composite likelihood's inherent property of being less sensitive to errors in individual calibration points, as demonstrated in simulations modeling various prior misspecifications.
摘要: Estimating divergence times from molecular sequence data is central to reconstructing the evolutionary history of lineages. Although Bayesian relaxed-clock methods provide a principled framework for incorporating fossil information, their dependence on repeated evaluations of the full phylogenetic likelihood makes them computationally demanding for large genomic datasets. Furthermore, because disagreements in divergence-time estimates often arise from uncertainty or error in fossil placement and prior specification, there is a need for methods that are both computationally efficient and robust to fossil-calibration uncertainty. In this study, we introduce fast and accurate alternatives based on the phylogenetic pairwise composite likelihood, presenting two adjusted pairwise likelihood (APW) formulations that employ asymptotic moment-matching weights to better approximate the behavior of the full likelihood within a Bayesian MCMC framework. Extensive simulations across diverse fossil-calibration scenarios show that APW methods produce node-age estimates comparable to those obtained from the full likelihood while offering greater robustness to fossil misplacement and prior misspecification, due to the reduced sensitivity of composite likelihoods to local calibration errors. Applied to a genome-scale dataset of modern birds, APW methods recover divergence time patterns consistent with recent studies, while reducing computational cost by more than an order of magnitude. Overall, our results demonstrate that adjusted pairwise likelihoods provide a calibration-robust and computationally efficient framework for Bayesian node dating, especially suited for large phylogenomic datasets and analyses in which fossil priors may be uncertain or imperfectly placed.