Paper List
-
Discovery of a Hematopoietic Manifold in scGPT Yields a Method for Extracting Performant Algorithms from Biological Foundation Model Internals
This work addresses the core challenge of extracting reusable, interpretable, and high-performance biological algorithms from the opaque internal repr...
-
MS2MetGAN: Latent-space adversarial training for metabolite–spectrum matching in MS/MS database search
This paper addresses the critical bottleneck in metabolite identification: the generation of high-quality negative training samples that are structura...
-
Toward Robust, Reproducible, and Widely Accessible Intracranial Language Brain-Computer Interfaces: A Comprehensive Review of Neural Mechanisms, Hardware, Algorithms, Evaluation, Clinical Pathways and Future Directions
This review addresses the core challenge of fragmented and heterogeneous evidence that hinders the clinical translation of intracranial language BCIs,...
-
Less Is More in Chemotherapy of Breast Cancer
通过纳入细胞周期时滞和竞争项,解决了现有肿瘤-免疫模型的过度简化问题,以定量比较化疗方案。
-
Fold-CP: A Context Parallelism Framework for Biomolecular Modeling
This paper addresses the critical bottleneck of GPU memory limitations that restrict AlphaFold 3-like models to processing only a few thousand residue...
-
Open Biomedical Knowledge Graphs at Scale: Construction, Federation, and AI Agent Access with Samyama Graph Database
This paper addresses the core pain point of fragmented biomedical data by constructing and federating large-scale, open knowledge graphs to enable sea...
-
Predictive Analytics for Foot Ulcers Using Time-Series Temperature and Pressure Data
This paper addresses the critical need for continuous, real-time monitoring of diabetic foot health by developing an unsupervised anomaly detection fr...
-
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, wh...
Training Dynamics of Learning 3D-Rotational Equivariance
Genentech Computational Sciences | New York University
30秒速读
IN SHORT: This work addresses the core dilemma of whether to use computationally expensive equivariant architectures or faster symmetry-agnostic models with data augmentation, by quantifying the speed and extent to which the latter learn 3D rotational symmetry.
核心创新
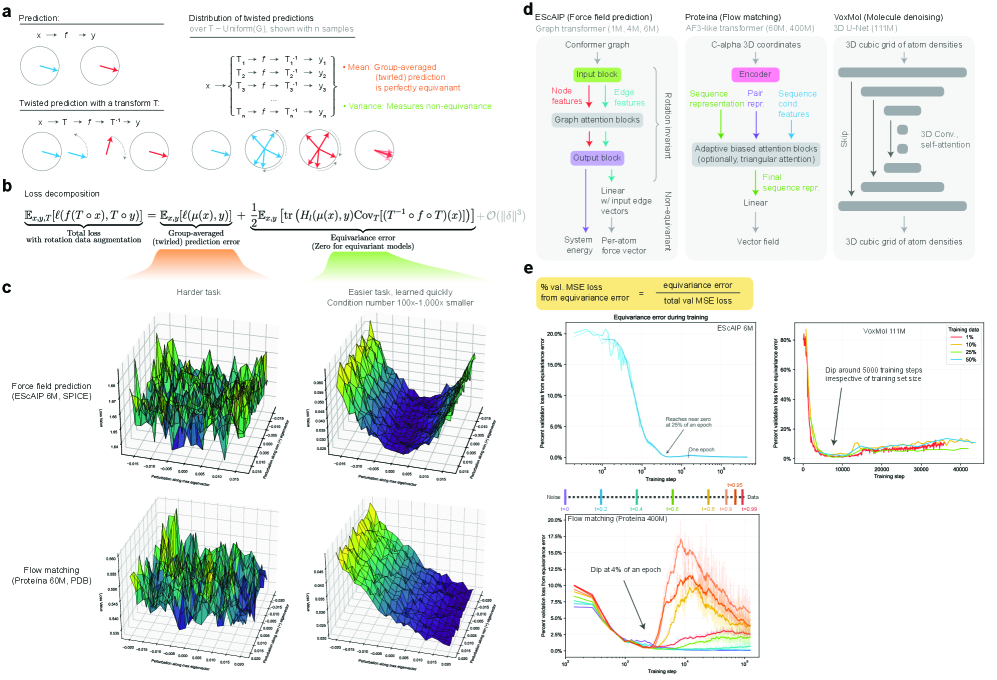
- Methodology Introduces a principled, generalizable framework to decompose total loss into a 'twirled prediction error' (ℒ_mean) and an 'equivariance error' (ℒ_equiv), enabling precise measurement of the percent of loss attributable to imperfect symmetry learning.
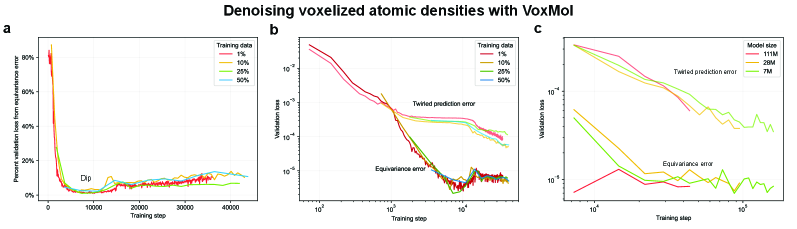
- Methodology Empirically demonstrates that models learning 3D-rotational equivariance via data augmentation achieve very low equivariance error (≤2% of total loss) remarkably quickly, within 1k-10k training steps, across diverse molecular tasks and model scales.
- Theory Provides theoretical and experimental evidence that learning equivariance is an easier task than the main prediction, characterized by a smoother and better-conditioned loss landscape (e.g., 1000x lower condition number for ℒ_equiv vs. ℒ_mean in force field prediction).
主要结论
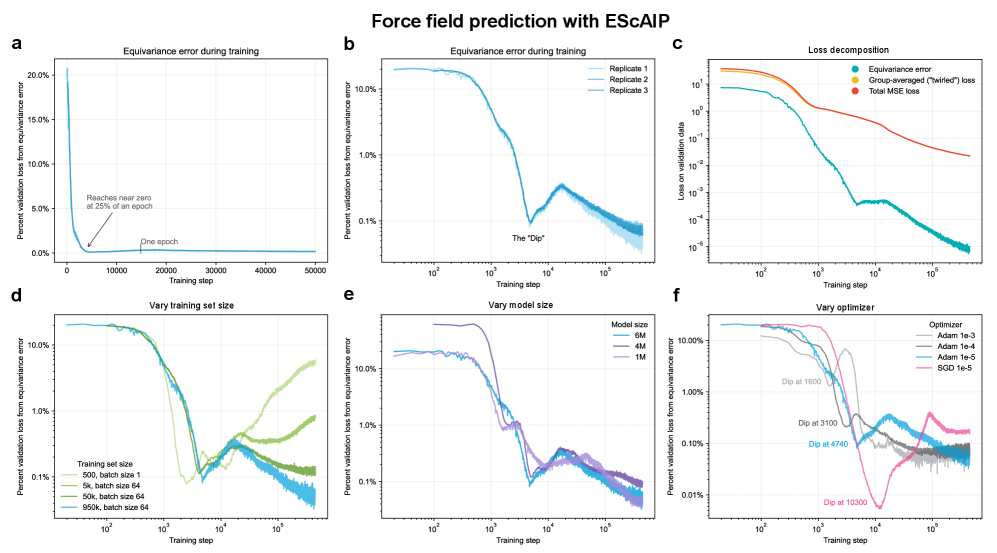
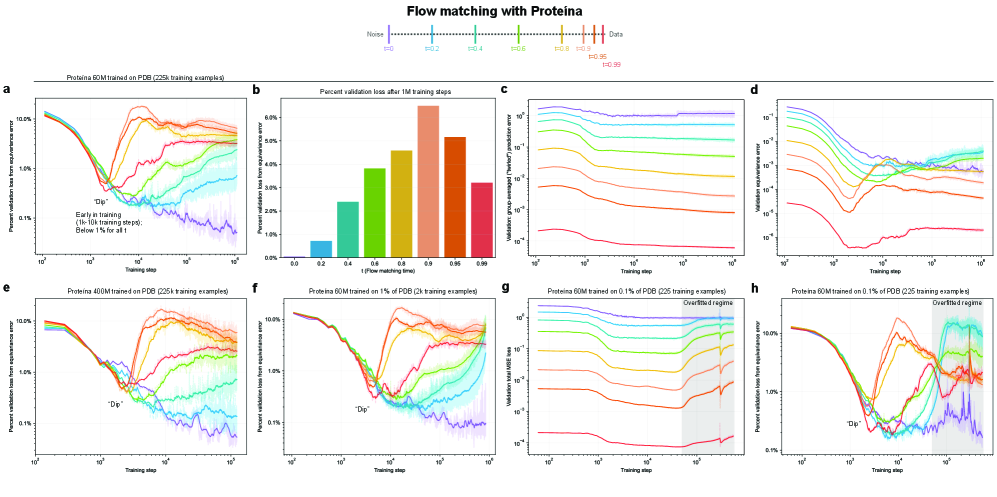
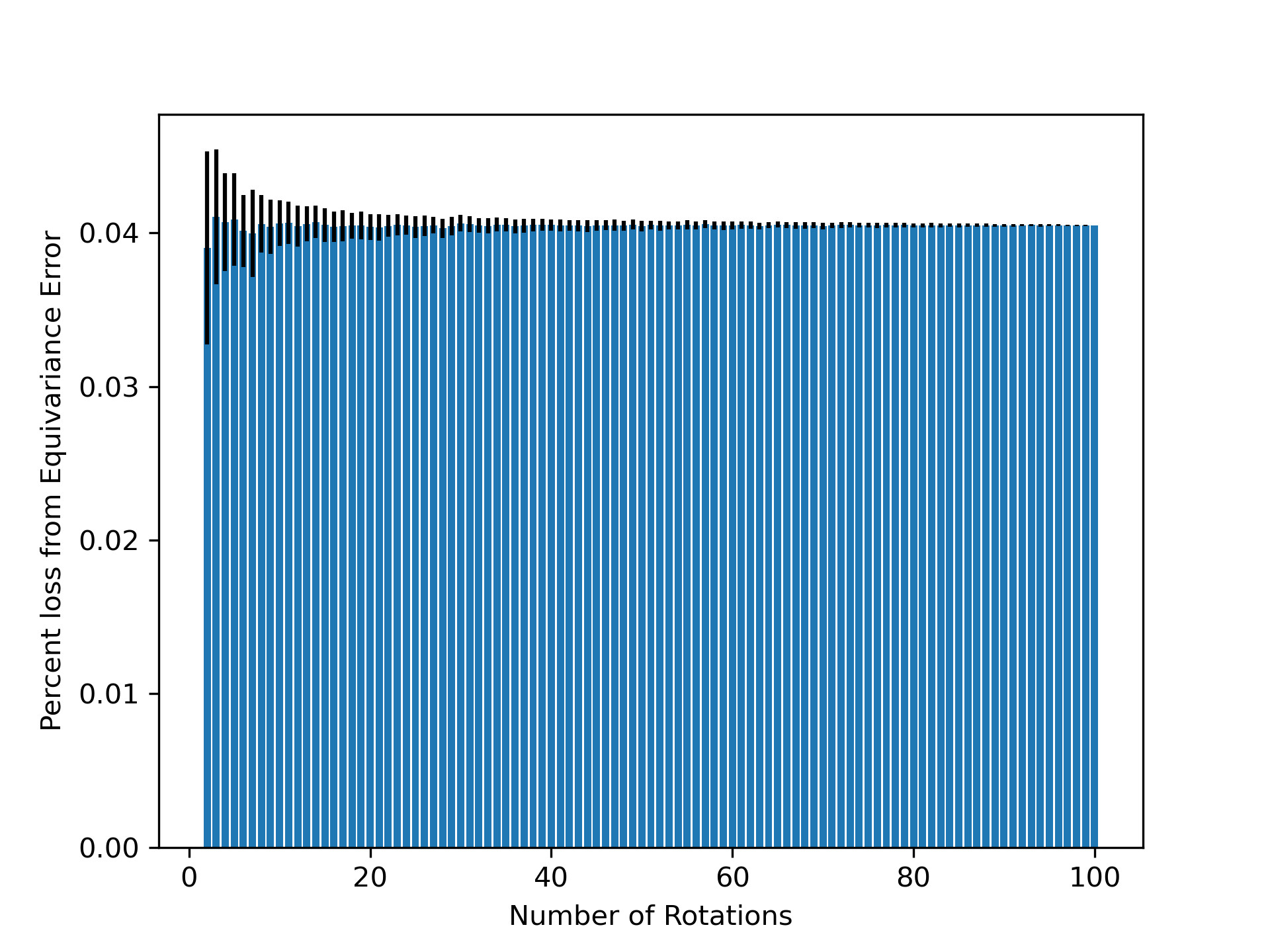
- Non-equivariant models with data augmentation learn 3D rotational equivariance rapidly and effectively, reducing the equivariance error component to ≤2% of the total validation loss within the first 1k-10k training steps.
- The loss penalty for imperfect equivariance (ℒ_equiv) is small throughout training for 3D rotations, meaning the primary trade-off is the 'efficiency gap' (slower training/inference of equivariant models) rather than a significant accuracy penalty.
- The speed of learning equivariance is robust to model size (1M to 400M parameters), dataset size (500 to 1M samples), and optimizer choice, indicating it is a fundamental property of the learning task landscape.
摘要: While data augmentation is widely used to train symmetry-agnostic models, it remains unclear how quickly and effectively they learn to respect symmetries. We investigate this by deriving a principled measure of equivariance error that, for convex losses, calculates the percent of total loss attributable to imperfections in learned symmetry. We focus our empirical investigation to 3D-rotation equivariance on high-dimensional molecular tasks (flow matching, force field prediction, denoising voxels) and find that models reduce equivariance error quickly to ≤2% held-out loss within 1k-10k training steps, a result robust to model and dataset size. This happens because learning 3D-rotational equivariance is an easier learning task, with a smoother and better-conditioned loss landscape, than the main prediction task. For 3D rotations, the loss penalty for non-equivariant models is small throughout training, so they may achieve lower test loss than equivariant models per GPU-hour unless the equivariant “efficiency gap” is narrowed. We also experimentally and theoretically investigate the relationships between relative equivariance error, learning gradients, and model parameters.