Paper List
-
The Effective Reproduction Number in the Kermack-McKendrick model with age of infection and reinfection
This paper addresses the challenge of accurately estimating the time-varying effective reproduction number ℛ(t) in epidemics by incorporating two crit...
-
Covering Relations in the Poset of Combinatorial Neural Codes
This work addresses the core challenge of navigating the complex poset structure of neural codes to systematically test the conjecture linking convex ...
-
Collective adsorption of pheromones at the water-air interface
This paper addresses the core challenge of understanding how amphiphilic pheromones, previously assumed to be transported in the gas phase, can be sta...
-
pHapCompass: Probabilistic Assembly and Uncertainty Quantification of Polyploid Haplotype Phase
This paper addresses the core challenge of accurately assembling polyploid haplotypes from sequencing data, where read assignment ambiguity and an exp...
-
Setting up for failure: automatic discovery of the neural mechanisms of cognitive errors
This paper addresses the core challenge of automating the discovery of biologically plausible recurrent neural network (RNN) dynamics that can replica...
-
Influence of Object Affordance on Action Language Understanding: Evidence from Dynamic Causal Modeling Analysis
This study addresses the core challenge of moving beyond correlational evidence to establish the *causal direction* and *temporal dynamics* of how obj...
-
Revealing stimulus-dependent dynamics through statistical complexity
This paper addresses the core challenge of detecting stimulus-specific patterns in neural population dynamics that remain hidden to traditional variab...
-
Exactly Solvable Population Model with Square-Root Growth Noise and Cell-Size Regulation
This paper addresses the fundamental gap in understanding how microscopic growth fluctuations, specifically those with size-dependent (square-root) no...
Mapping of Lesion Images to Somatic Mutations
University of Illinois at Chicago | University of Texas MD Anderson Cancer Center
30秒速读
IN SHORT: This paper addresses the critical bottleneck of delayed genetic analysis in cancer diagnosis by predicting a patient's full somatic mutation profile directly from medical lesion images, enabling earlier targeted treatment decisions.
核心创新
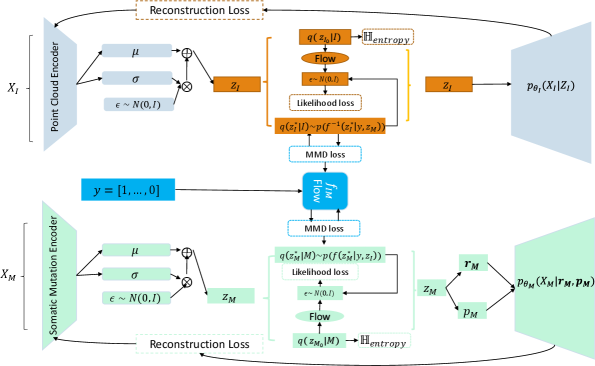
- Methodology Proposes LLOST, a novel architecture with dual VAEs and a separate, cancer-type-conditioned shared latent space, coupled with domain-specific conditional Normalizing Flow priors to handle heterogeneous data distributions.
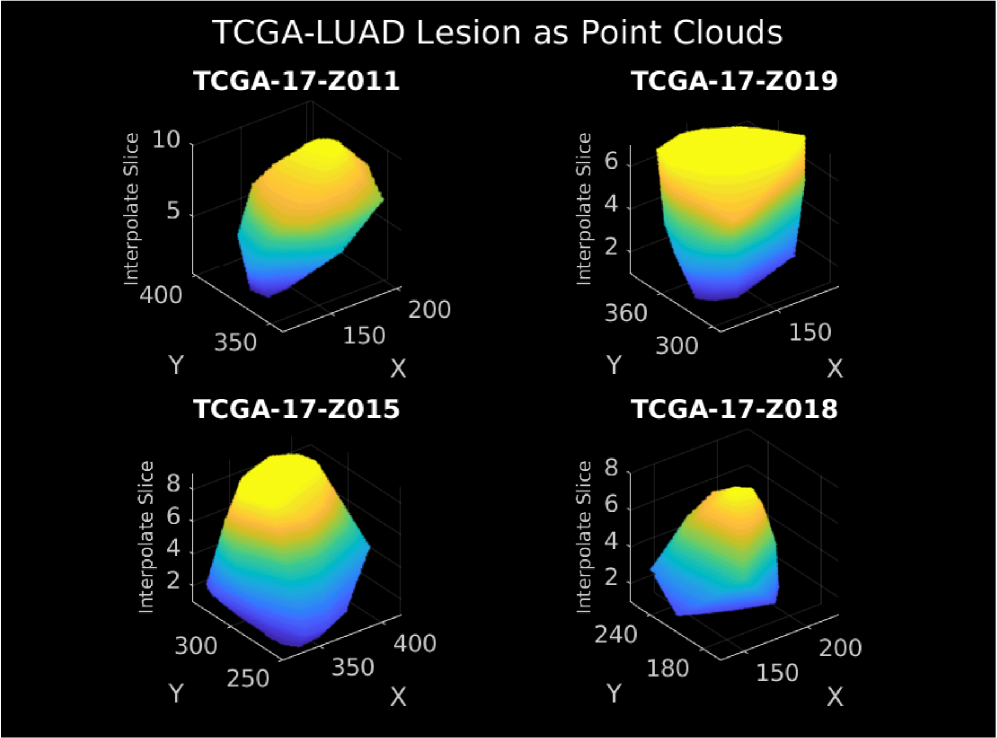
- Methodology Introduces a modality-invariant point cloud representation for lesion images, overcoming challenges of multi-slice, multi-modal (CT/MRI) medical imaging data.
- Methodology Employs a Negative-Binomial likelihood within the mutation VAE to effectively model the high-dimensional, sparse, and discrete nature of somatic mutation count data.
主要结论
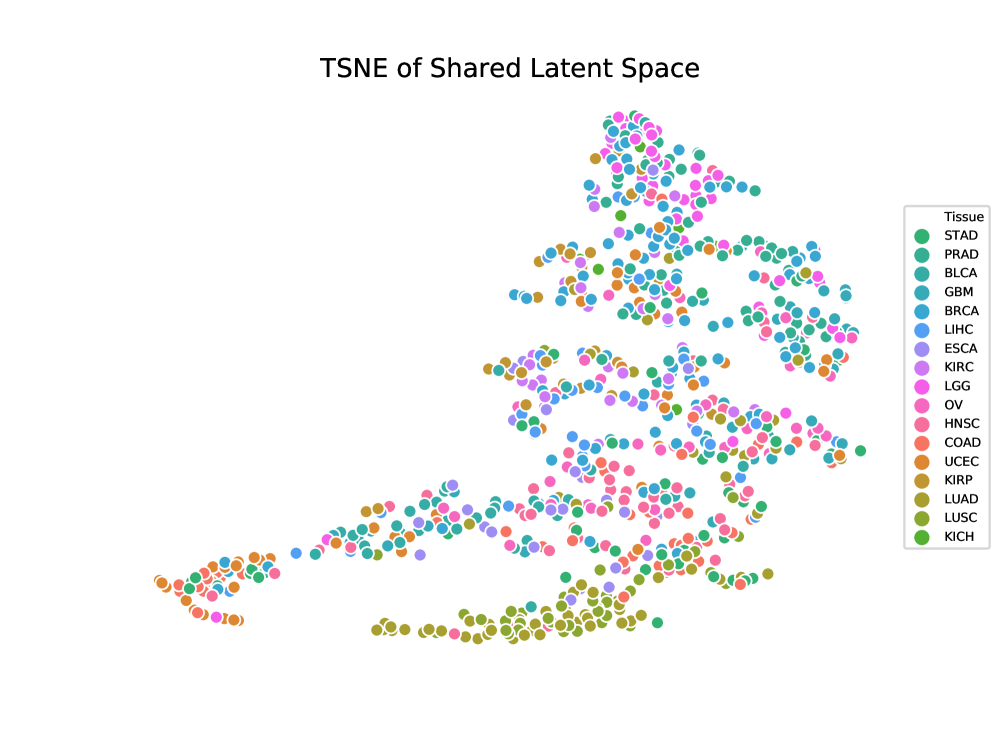
- LLOST successfully learns a shared latent representation between lesion point clouds and somatic mutation counts, capturing cancer-type-specific patterns across these disparate domains.
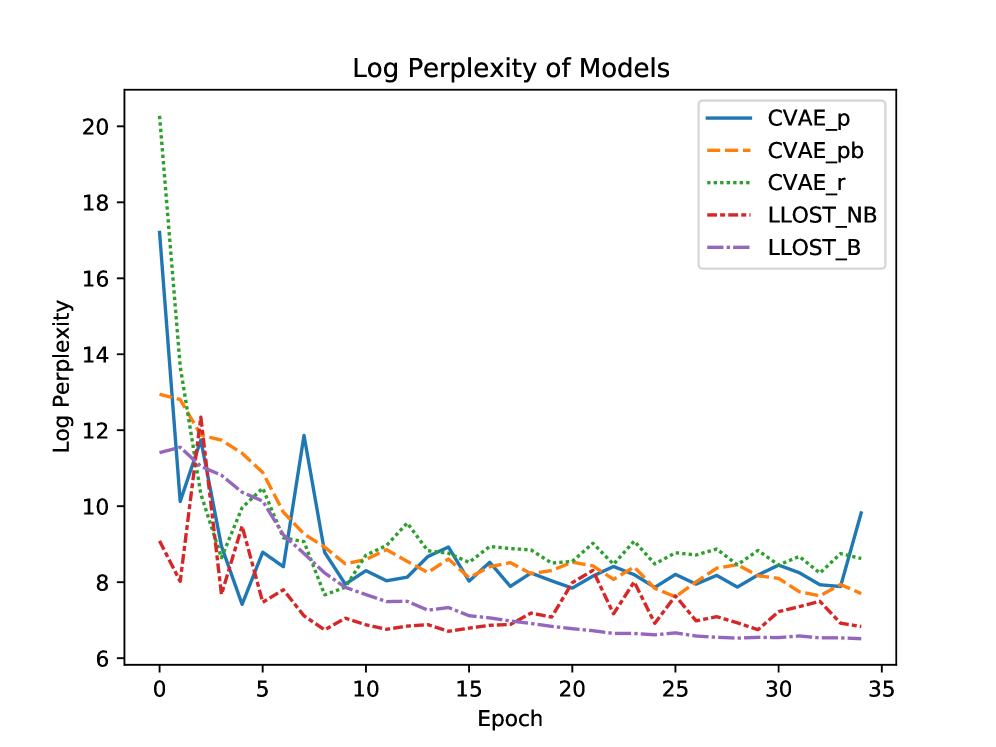
- The model demonstrates predictive capability for both mutation occurrence (binary prediction) and mutation counts, validated on a dataset of 1342 patients across 18 cancer types from TCGA/TCIA.
- The use of conditional Normalizing Flow priors and a separate shared latent space allows the model to account for and bridge the complex, distinct distributions of imaging and genomic data.
摘要: Medical imaging is a critical initial tool used by clinicians to determine a patient’s cancer diagnosis, allowing for faster intervention and more reliable patient prognosis. At subsequent stages of patient diagnosis, genetic information is extracted to help select specific patient treatment options. As the efficacy of cancer treatment often relies on early diagnosis and treatment, we build a deep latent variable model to determine patients’ somatic mutation profiles based on their corresponding medical images. We first introduce a point cloud representation of lesions images to allow for invariance to the imaging modality. We then propose, LLOST, a model with dual variational autoencoders coupled together by a separate shared latent space that unifies features from the lesion point clouds and counts of distinct somatic mutations. Therefore our model consists of three latent space, each of which is learned with a conditional normalizing flow prior to account for the diverse distributions of each domain. We conduct qualitative and quantitative experiments on de-identified medical images from The Cancer Imaging Archive and the corresponding somatic mutations from the Pan Cancer dataset of The Cancer Genomic Archive. We show the model’s predictive performance on the counts of specific mutations as well as it’s ability to accurately predict the occurrence of mutations. In particular, shared patterns between the imaging and somatic mutation domain that reflect cancer type. We conclude with a remark on how to improve the model and possible future avenues of research to include other genetic domains.