Paper List
-
Formation of Artificial Neural Assemblies by Biologically Plausible Inhibition Mechanisms
This work addresses the core limitation of the Assembly Calculus model—its fixed-size, biologically implausible k-WTA selection process—by introducing...
-
How to make the most of your masked language model for protein engineering
This paper addresses the critical bottleneck of efficiently sampling high-quality, diverse protein sequences from Masked Language Models (MLMs) for pr...
-
Module control in youth symptom networks across COVID-19
This paper addresses the core challenge of distinguishing whether a prolonged societal stressor (COVID-19) fundamentally reorganizes the architecture ...
-
JEDI: Jointly Embedded Inference of Neural Dynamics
This paper addresses the core challenge of inferring context-dependent neural dynamics from noisy, high-dimensional recordings using a single unified ...
-
ATP Level and Phosphorylation Free Energy Regulate Trigger-Wave Speed and Critical Nucleus Size in Cellular Biochemical Systems
This work addresses the core challenge of quantitatively predicting how the cellular energy state (ATP level and phosphorylation free energy) governs ...
-
Packaging Jupyter notebooks as installable desktop apps using LabConstrictor
This paper addresses the core pain point of ensuring Jupyter notebook reproducibility and accessibility across different computing environments, parti...
-
SNPgen: Phenotype-Supervised Genotype Representation and Synthetic Data Generation via Latent Diffusion
This paper addresses the core challenge of generating privacy-preserving synthetic genotype data that maintains both statistical fidelity and downstre...
-
Continuous Diffusion Transformers for Designing Synthetic Regulatory Elements
This paper addresses the challenge of efficiently generating novel, cell-type-specific regulatory DNA sequences with high predicted activity while min...
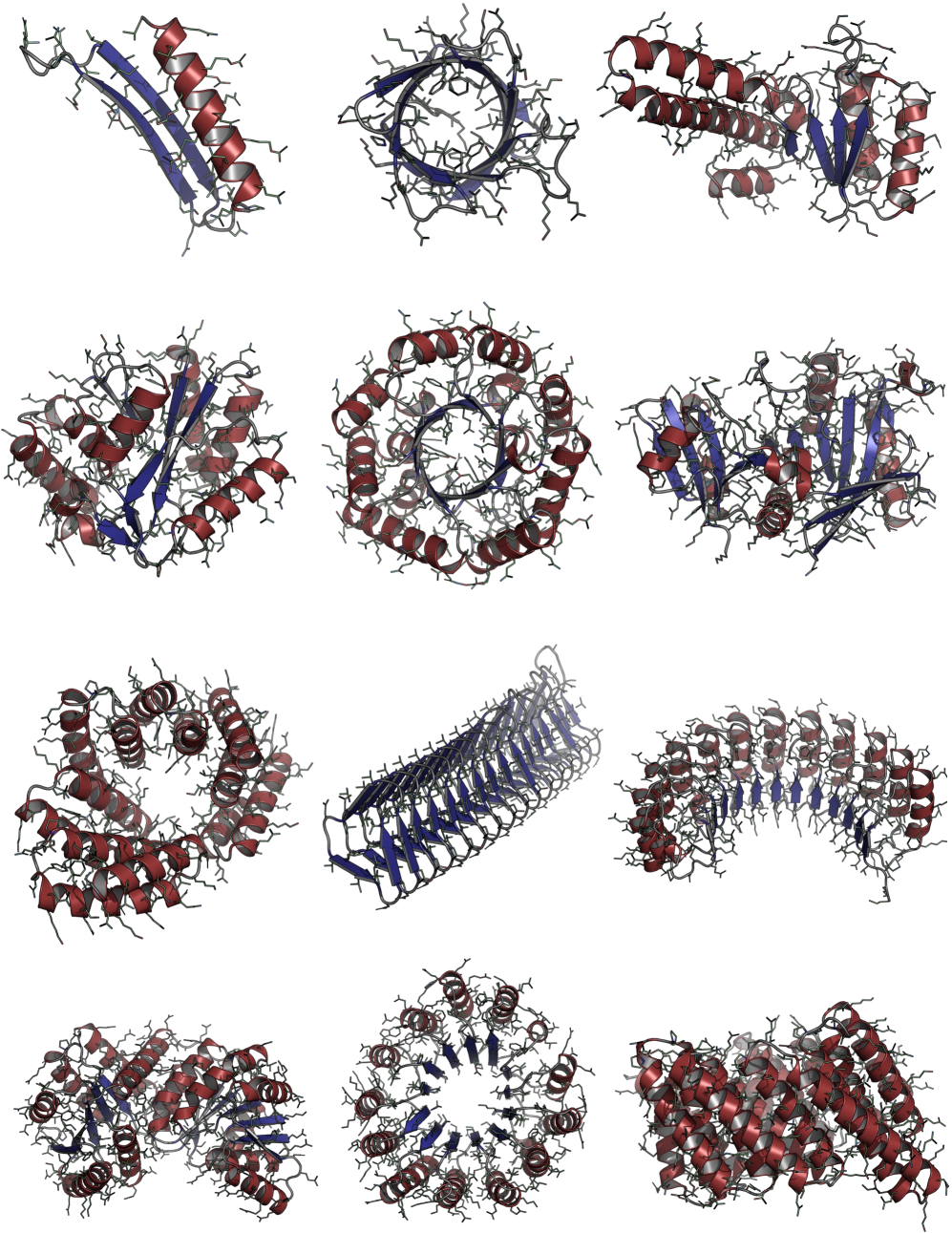
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
NVIDIA | Mila - Quebec AI Institute | Université de Montréal | HEC Montréal | CIFAR AI Chair
30秒速读
IN SHORT: This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the performance of fully atomistic protein generative models.
核心创新
- Methodology Introduces a novel high-quality synthetic dataset (D_SYN-ours, ~0.46M samples) by leveraging ProteinMPNN for sequence generation and ESMFold for refolding, ensuring aligned and recoverable sequence-structure pairs.
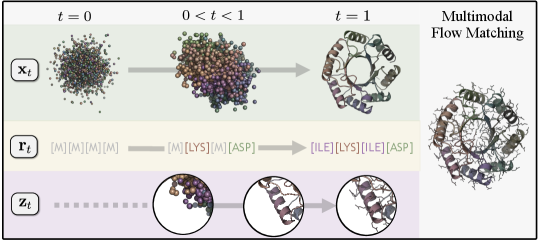
- Methodology Proposes Proteína-Atomística, a unified multi-modal flow-based framework that jointly models the distribution of Cα backbone atoms, discrete amino acid sequences, and non-Cα side-chain atoms in explicit observable space without latent variables.
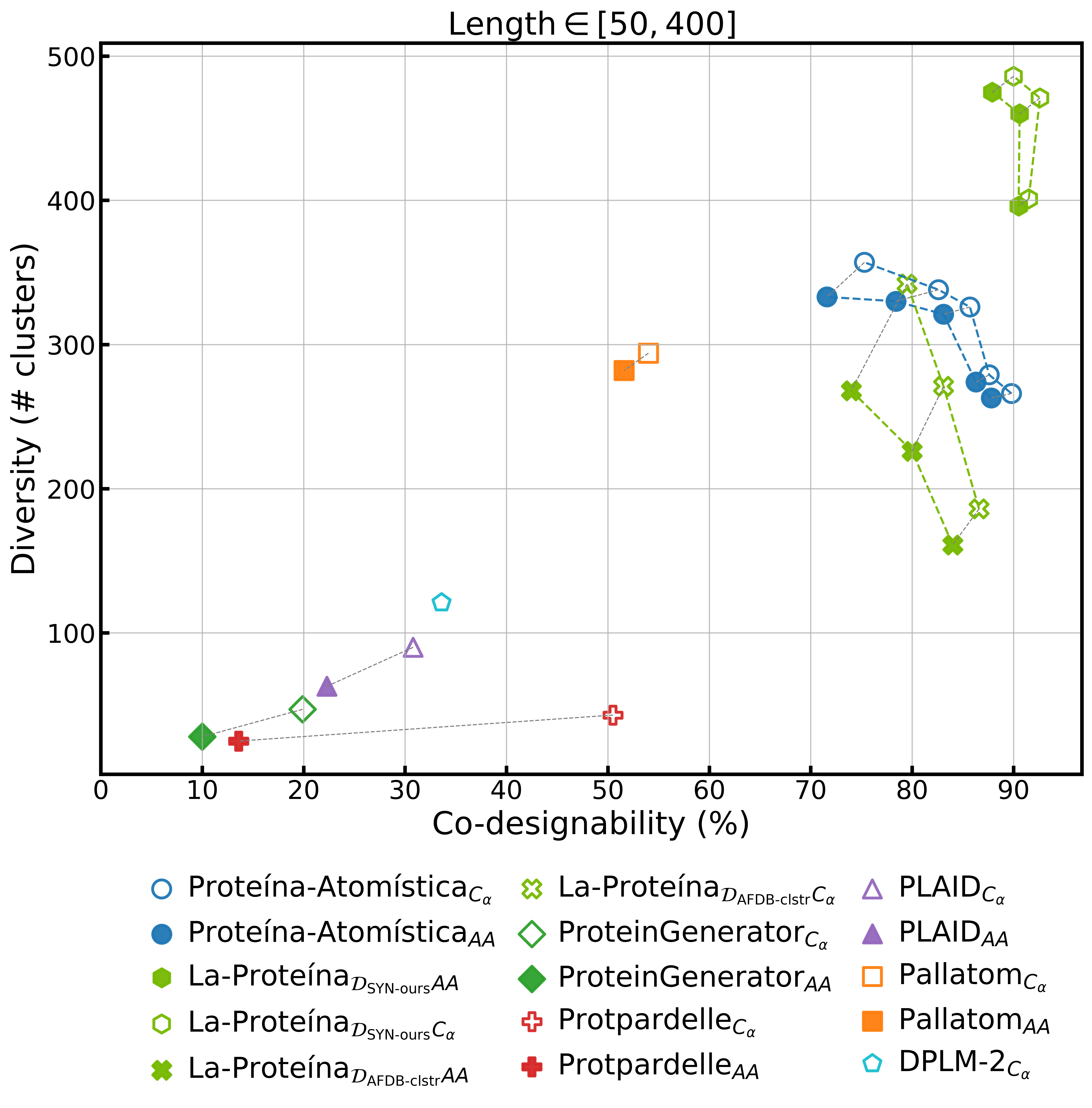
- Biology Demonstrates that consistent synthetic sequences are critical for unlocking structural diversity, with retrained La-Proteína achieving +54% structural diversity and +27% co-designability, and Proteína-Atomística achieving +73% structural diversity and +5% co-designability.
主要结论
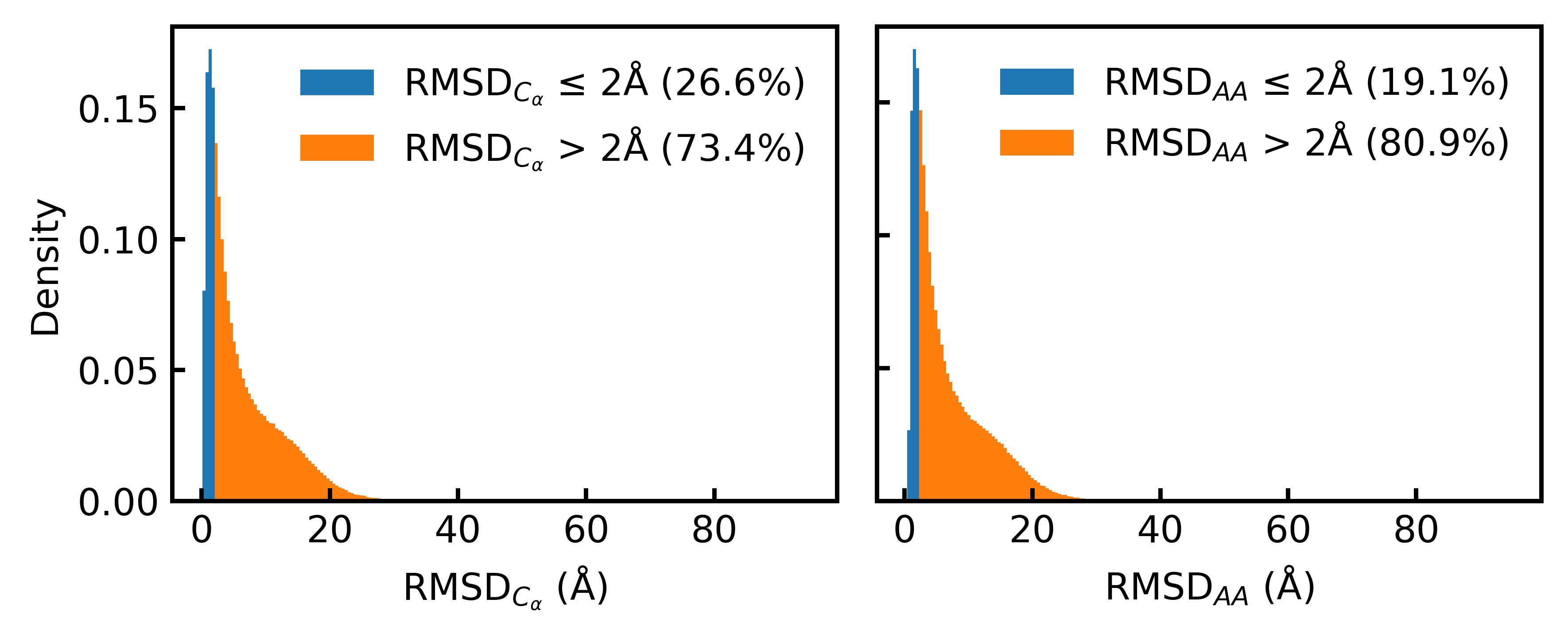
- Only 19.1% of the Foldseek-clustered AFDB dataset (D_AFDB-clstr) meets the standard 2Å all-atom RMSD co-designability threshold when refolded with ESMFold, revealing severe sequence-structure misalignment.
- Training on the new aligned dataset D_SYN-ours boosts La-Proteína's performance by +54% in structural diversity and +27% in co-designability, setting a new state-of-the-art.
- The proposed Proteína-Atomística framework, when trained on D_SYN-ours, shows a dramatic +73% improvement in structural diversity and a +5% improvement in co-designability, validating the dataset's broad utility.
摘要: High-quality training datasets are crucial for the development of effective protein design models, but existing synthetic datasets often include unfavorable sequence-structure pairs, impairing generative model performance. We leverage ProteinMPNN, whose sequences are experimentally favorable as well as amenable to folding, together with structure prediction models to align high-quality synthetic structures with recoverable synthetic sequences. In that way, we create a new dataset designed specifically for training expressive, fully atomistic protein generators. By retraining La-Proteína, which models discrete residue type and side chain structure in a continuous latent space, on this dataset, we achieve new state-of-the-art results, with improvements of +54% in structural diversity and +27% in co-designability. To validate the broad utility of our approach, we further introduce Proteína-Atomística, a unified flow-based framework that jointly learns the distribution of protein backbone structure, discrete sequences, and atomistic side chains without latent variables. We again find that training on our new sequence-structure data dramatically boosts benchmark performance, improving Proteína-Atomística’s structural diversity by +73% and co-designability by +5%. Our work highlights the critical importance of aligned sequence-structure data for training high-performance de novo protein design models. All data will be publicly released.