Paper List
-
Mapping of Lesion Images to Somatic Mutations
This paper addresses the critical bottleneck of delayed genetic analysis in cancer diagnosis by predicting a patient's full somatic mutation profile d...
-
Reinventing Clinical Dialogue: Agentic Paradigms for LLM‑Enabled Healthcare Communication
This paper addresses the core challenge of transforming reactive, stateless LLMs into autonomous, reliable clinical dialogue agents capable of longitu...
-
Binary Latent Protein Fitness Landscapes for Quantum Annealing Optimization
通过将序列映射到二元潜在空间进行基于QUBO的适应度优化,桥接蛋白质表示学习和组合优化。
-
Controlling Fish Schools via Reinforcement Learning of Virtual Fish Movement
证明了无模型强化学习可以利用虚拟视觉刺激有效引导鱼群,克服了缺乏精确行为模型的问题。
EnzyCLIP: A Cross-Attention Dual Encoder Framework with Contrastive Learning for Predicting Enzyme Kinetic Constants
Vellore Institute of Technology | BIT (Department of Computer Science) | BIT (Department of Bioengineering and Biotechnology)
30秒速读
IN SHORT: This paper addresses the core challenge of jointly predicting enzyme kinetic parameters (Kcat and Km) by modeling dynamic enzyme-substrate interactions through a multimodal contrastive learning framework.
核心创新
- Methodology Proposes a CLIP-inspired dual-encoder architecture with bidirectional cross-attention that dynamically models enzyme-substrate interactions, overcoming the limitation of separate processing in existing methods.
- Methodology Integrates contrastive learning (InfoNCE loss) with multi-task regression (Huber loss) to learn aligned multimodal representations while jointly predicting both Kcat and Km parameters.
- Biology Addresses the critical gap in existing literature that typically focuses on single parameter prediction (mainly Kcat) by providing a unified framework for joint prediction of both fundamental kinetic constants.
主要结论
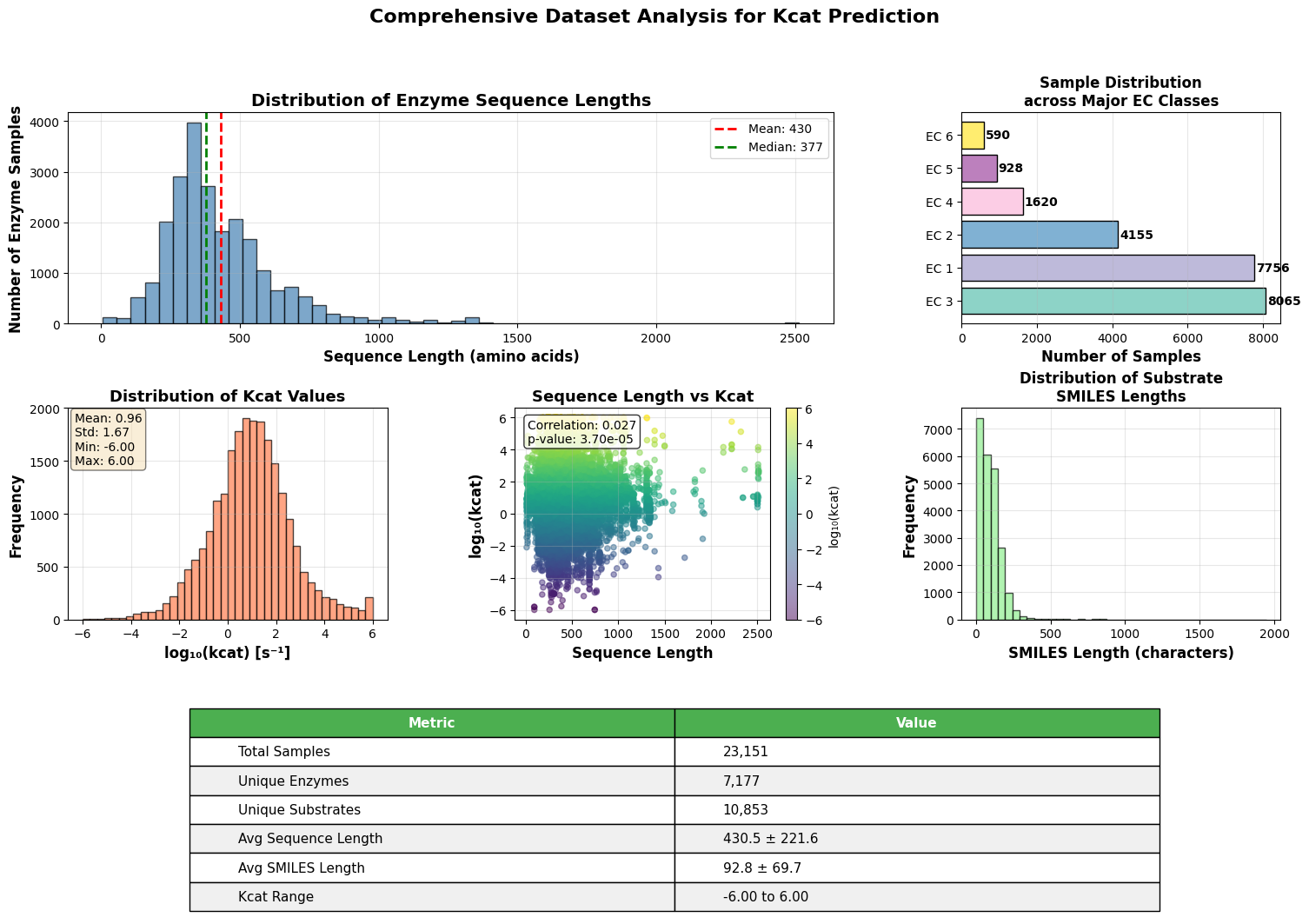
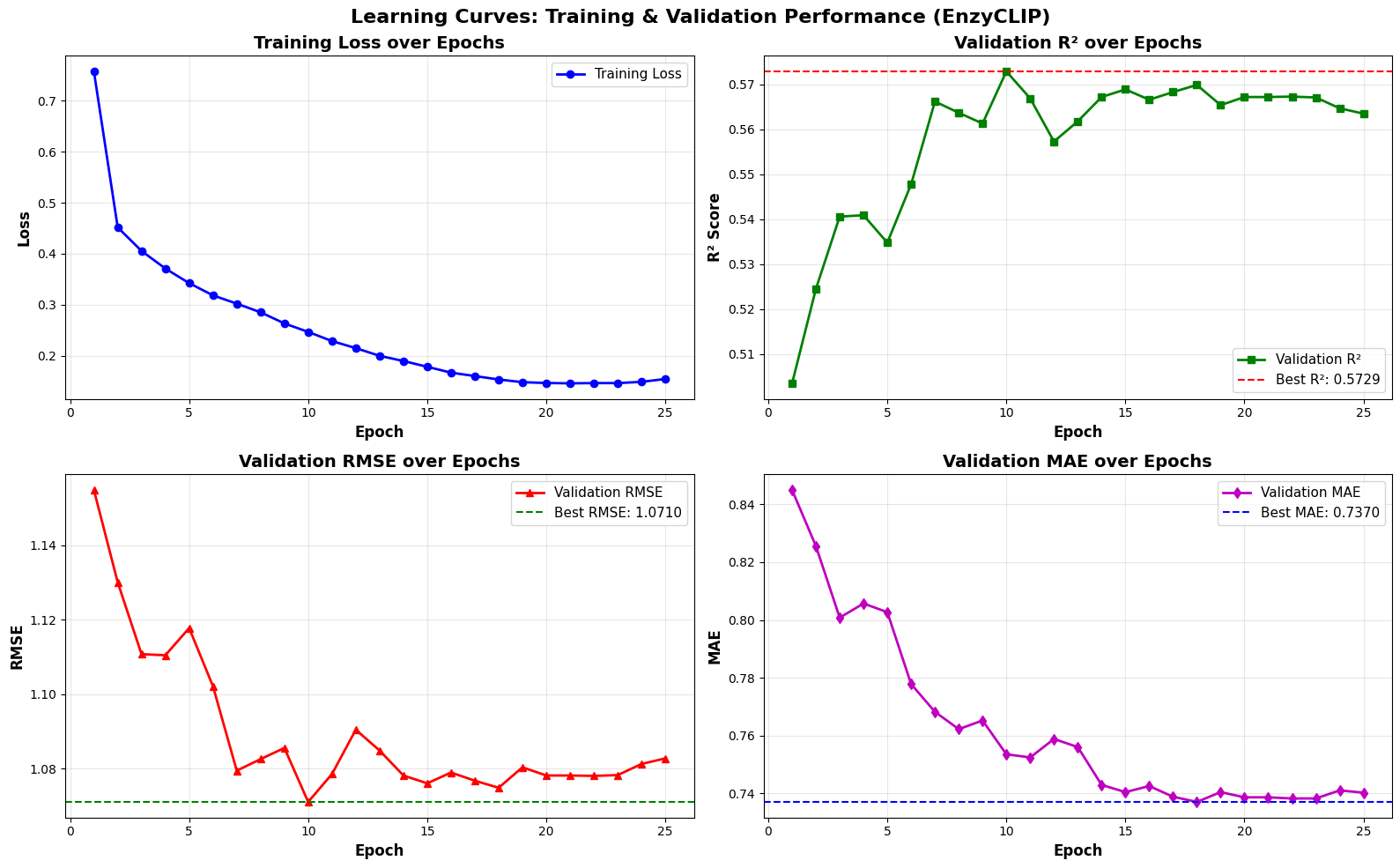
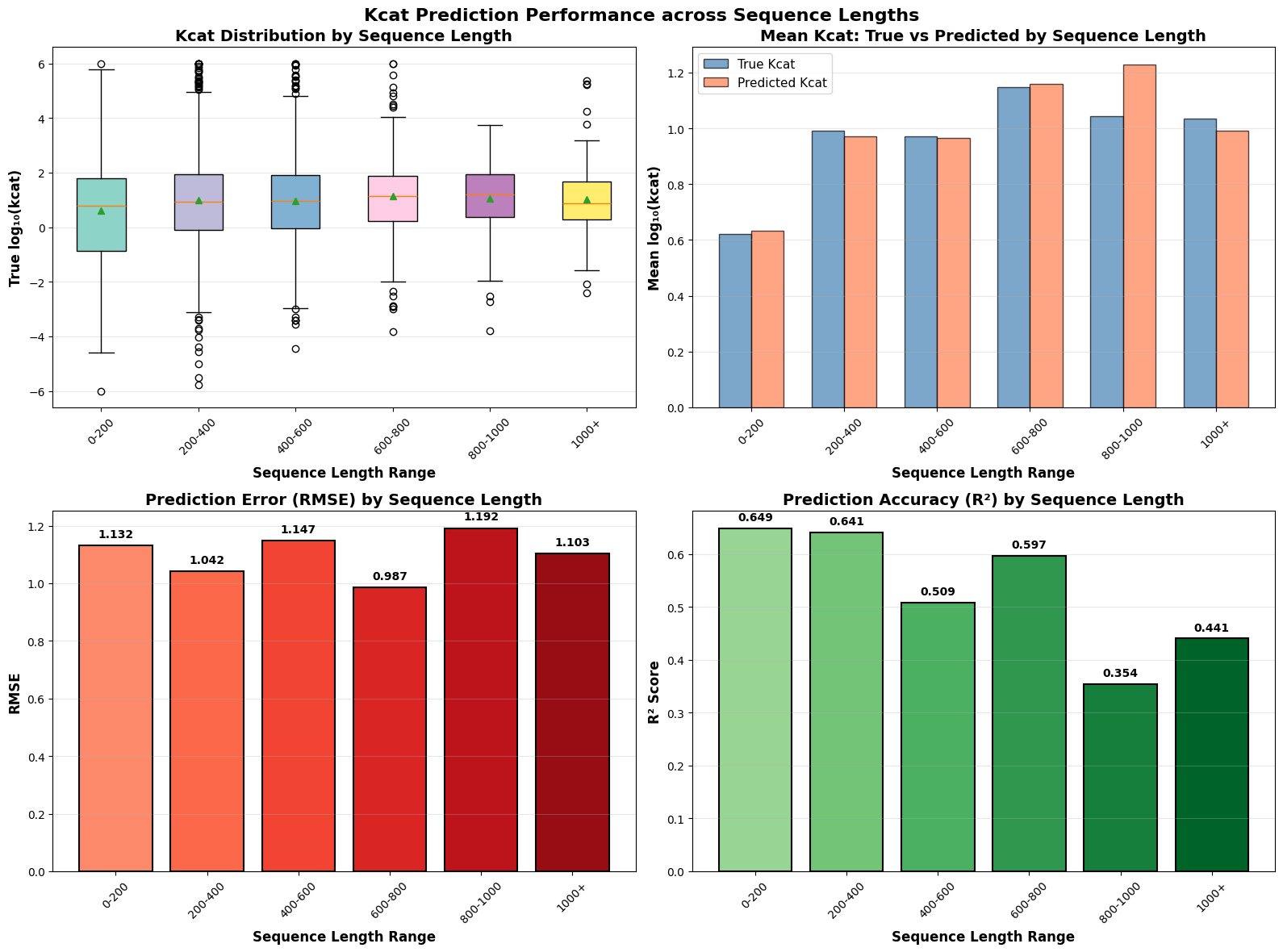
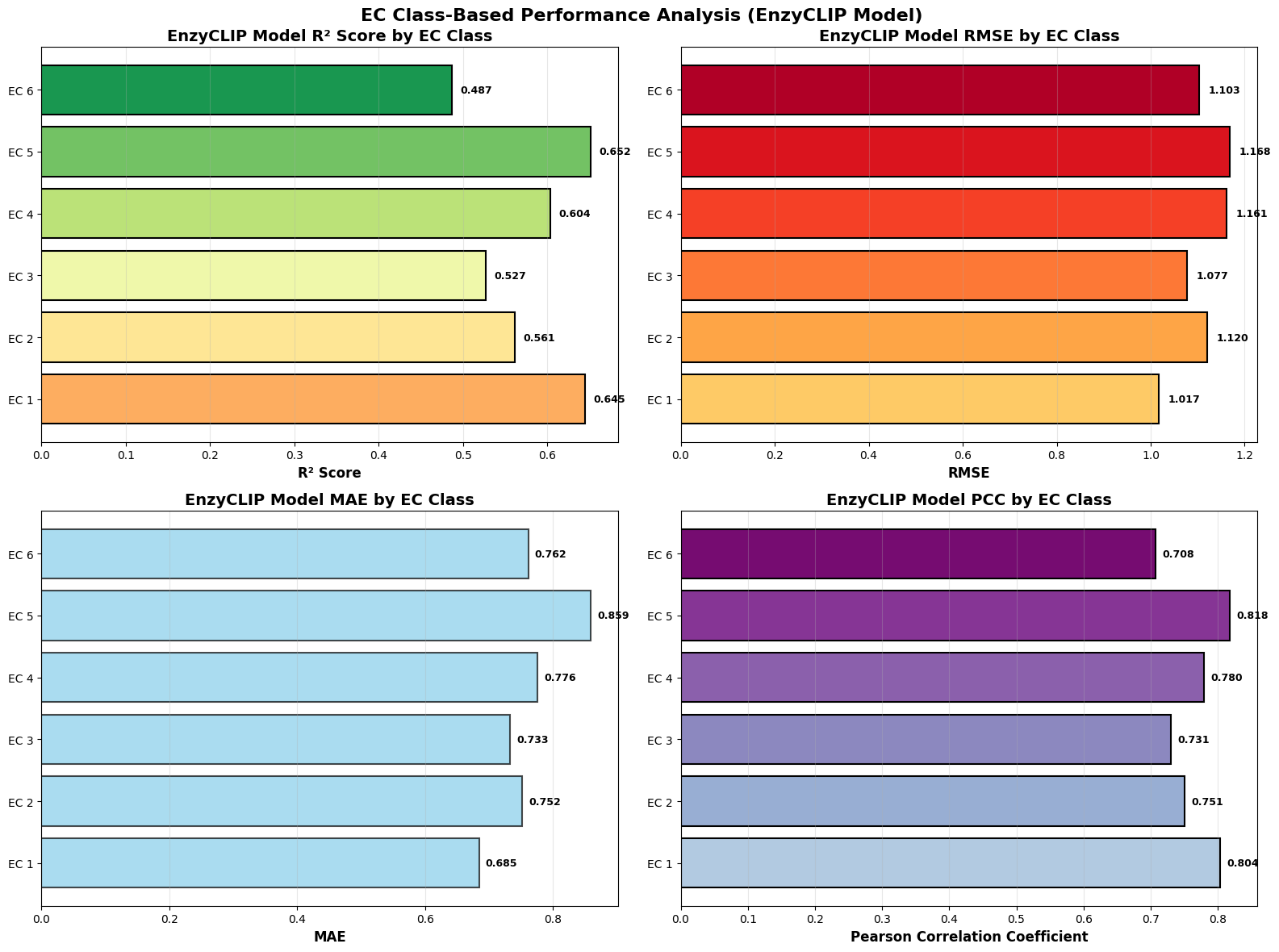
- EnzyCLIP achieves competitive baseline performance with R² scores of 0.593 for Kcat and 0.607 for Km prediction on the CatPred-DB dataset containing 23,151 Kcat and 41,174 Km measurements.
- The integration of contrastive learning with cross-attention mechanisms enables the model to capture biochemical relationships and substrate preferences even for unseen enzyme-substrate pairs.
- XGBoost ensemble methods applied to learned embeddings further improved Km prediction performance to R² = 0.61 while maintaining robust Kcat prediction capabilities.
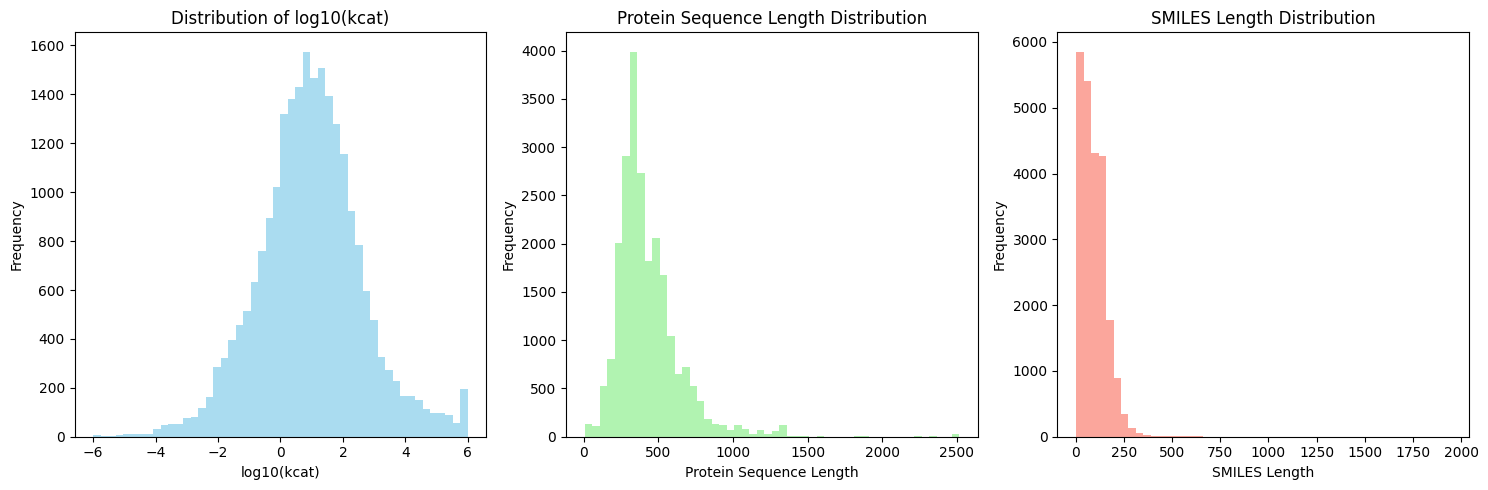
摘要: Accurate prediction of enzyme kinetic parameters is crucial for drug discovery, metabolic engineering, and synthetic biology applications. Current computational approaches face limitations in capturing complex enzyme–substrate interactions and often focus on single parameters while neglecting the joint prediction of catalytic turnover numbers (Kcat) and Michaelis–Menten constants (Km). We present EnzyCLIP, a novel dual-encoder framework that leverages contrastive learning and cross-attention mechanisms to predict enzyme kinetic parameters from protein sequences and substrate molecular structures. Our approach integrates ESM-2 protein language model embeddings with ChemBERTa chemical representations through a CLIP-inspired architecture enhanced with bidirectional cross-attention for dynamic enzyme–substrate interaction modeling. EnzyCLIP combines InfoNCE contrastive loss with Huber regression loss to learn aligned multimodal representations while predicting log10-transformed kinetic parameters. EnzyCLIP is trained on the CatPred-DB database containing 23,151 Kcat and 41,174 Km experimentally validated measurements, and achieved competitive baseline performance with R2 scores of 0.593 for Kcat and 0.607 for Km prediction. XGBoost ensemble methods on learned embeddings further improved Km prediction (R2 = 0.61) while maintaining robust Kcat performance.